A0A0H3C5R2
multicom
A0A0H3C5R2
full_length
A0A0H3C5R2
Results of Structure Prediction for Target Name: A0A0H3C5R2 ( Click  )
)
Domain Boundary prediction ( View  )
)

>A0A0H3C5R2: 1-179
| 1-60: |
M | R | R | P | L | P | T | P | E | E | A | I | A | I | L | R | S | K | R | T | K | P | Q | R | R | P | P | P | P | A | G | K | N | L | A | P | L | L | K | D | L | E | D | R | F | G | K | G | P | A | A | L | Q | A | R | W | K | E | I | V |
| 61-119: |
G | D | T | L | A | R | R | T | E | P | V | R | I | I | K | G | R | N | G | E | G | G | A | L | E | L | R | V | D | G | P | V | A | S | L | I | Q | H | Q | A | P | Q | I | T | A | R | L | D | M | L | L | G | K | G | V | V | T | R | L | R |
| 121-179: |
I | V | Q | G | P | V | K | A | P | A | A | A | P | S | T | R | L | R | R | K | P | P | L | D | A | A | L | E | K | Q | L | A | D | S | L | A | E | Q | P | D | G | A | L | K | Q | A | L | L | K | L | G | R | G | V | L | S | S | E | R |
| 1-60: |
C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | C | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H |
| 61-119: |
C | H | H | H | H | H | H | C | C | C | C | E | E | E | C | C | C | C | C | C | C | C | E | E | E | E | E | E | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | H | H | H | H | H | H | E | E |
| 121-179: |
E | E | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | C | C |
|
| | H(Helix): 100(55.87%) | E(Strand): 13(7.26%) | C(Coil): 66(36.87%) |
| 1-60: |
E | E | E | E | E | E | E | E | E | E | B | B | E | B | B | E | E | E | E | E | E | E | E | E | E | E | B | E | E | B | B | E | B | B | E | E | B | B | E | E | B | B | E | E | E | B | B | B | E | B | E | B | B | E | E | B | E | E | B | B |
| 61-119: |
B | E | E | B | B | E | B | B | E | B | E | E | B | E | E | E | E | E | E | E | E | B | B | B | B | B | B | B | E | B | B | B | B | B | B | B | B | B | E | B | E | E | B | B | E | B | B | B | E | B | B | B | B | E | B | B | E | E | B | E |
| 121-179: |
B | E | B | E | E | B | E | E | E | E | E | E | E | E | E | E | E | E | E | E | E | E | B | E | E | E | E | E | E | E | B | E | E | B | B | E | E | B | E | E | E | E | B | E | E | B | B | E | E | B | B | E | B | B | B | E | E | E | E |
|
| | e(Exposed): 105(58.66%) | b(Buried): 74(41.34%) |
| 1-60: |
T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 61-119: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 121-179: |
N | N | N | N | N | N | N | N | N | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | T | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | T | T | T |
|
| | N(Normal): 132(73.74%) | T(Disorder): 47(26.26%) |
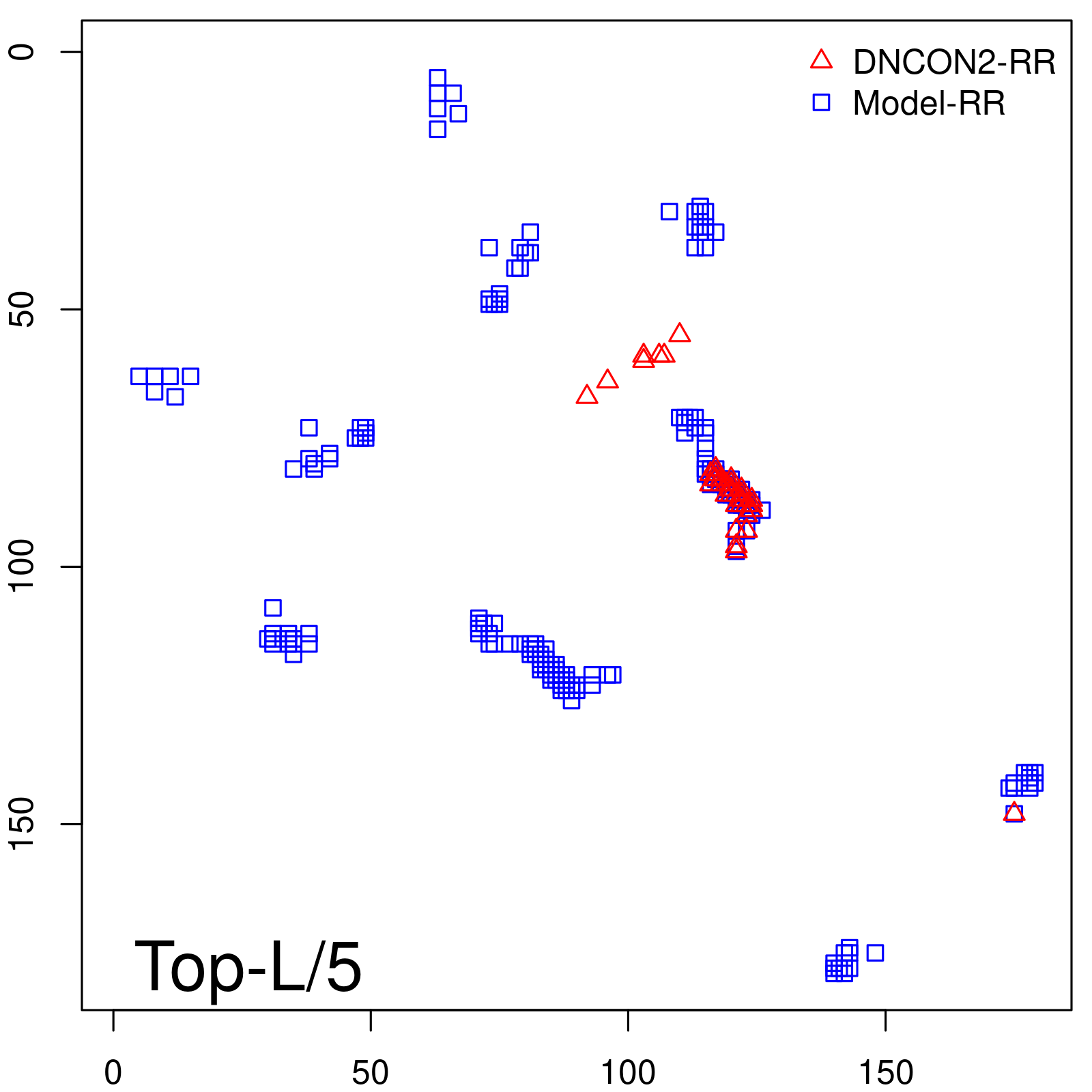
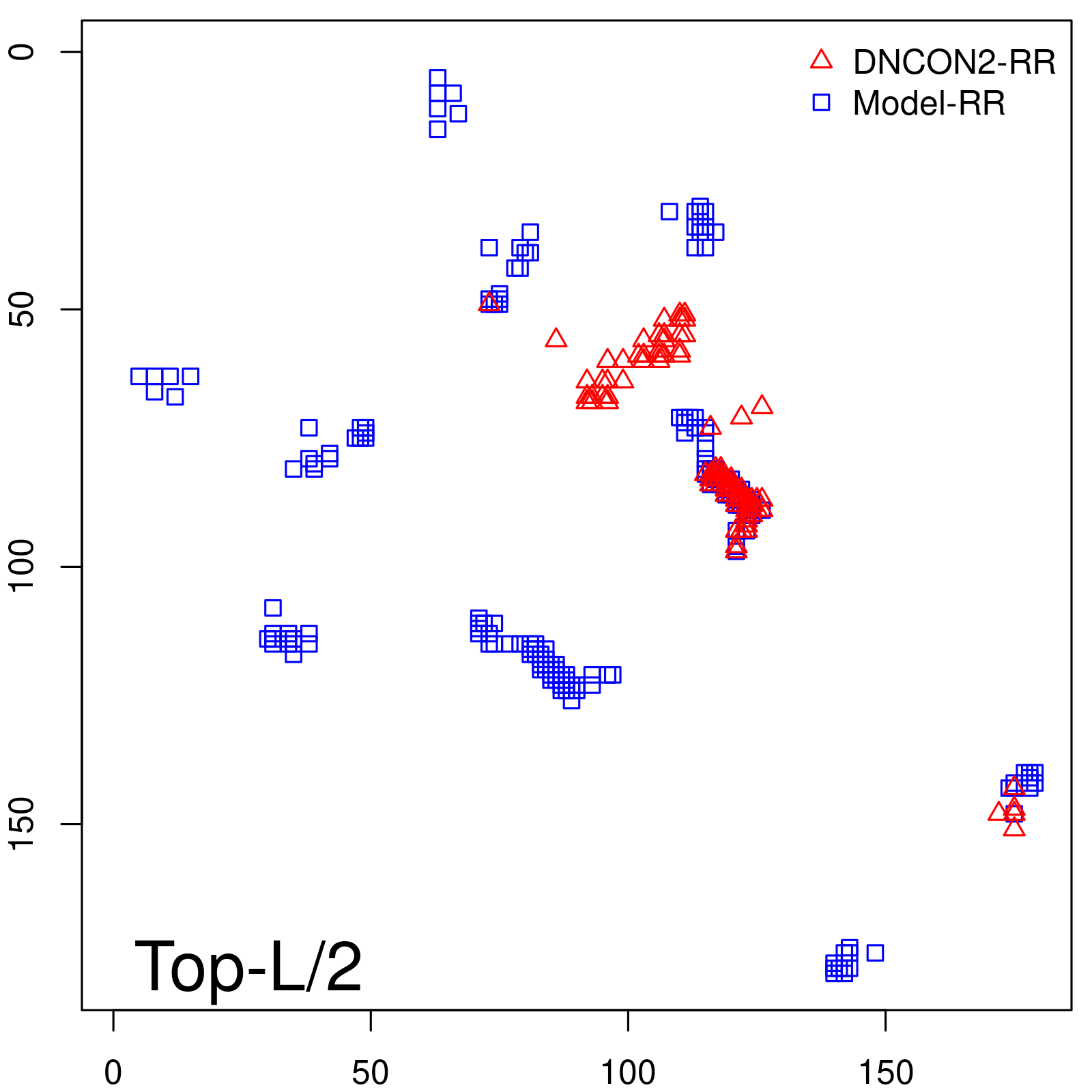
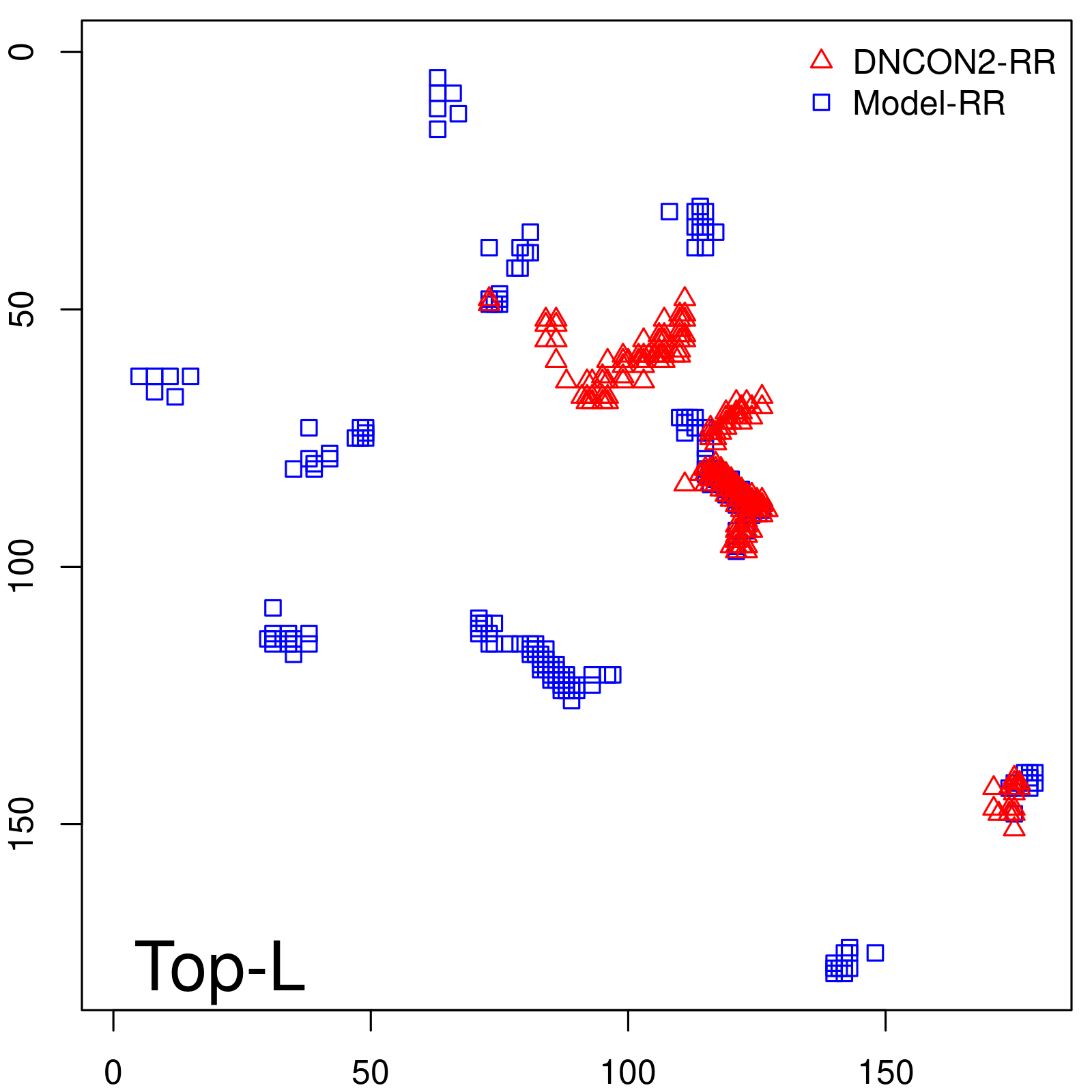
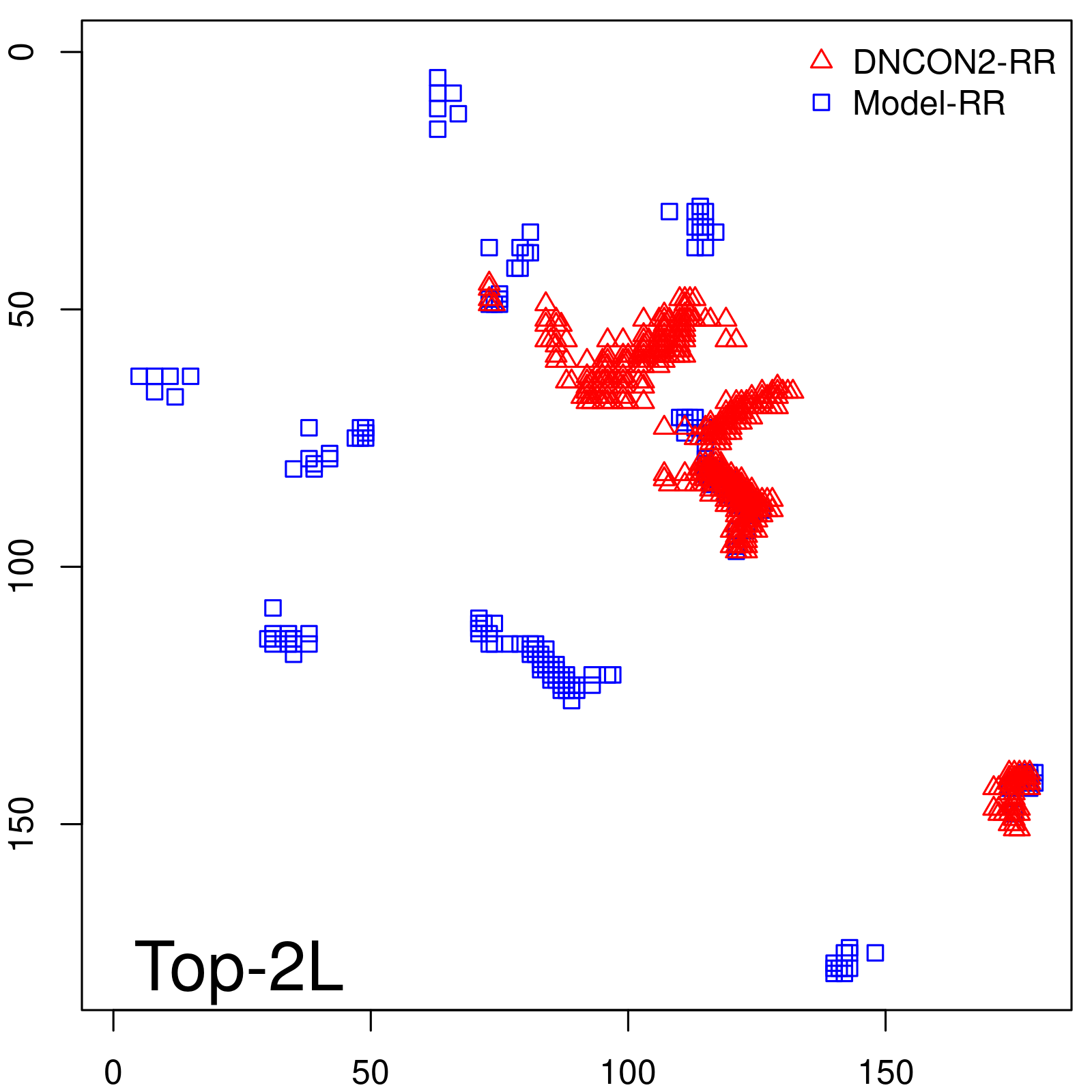
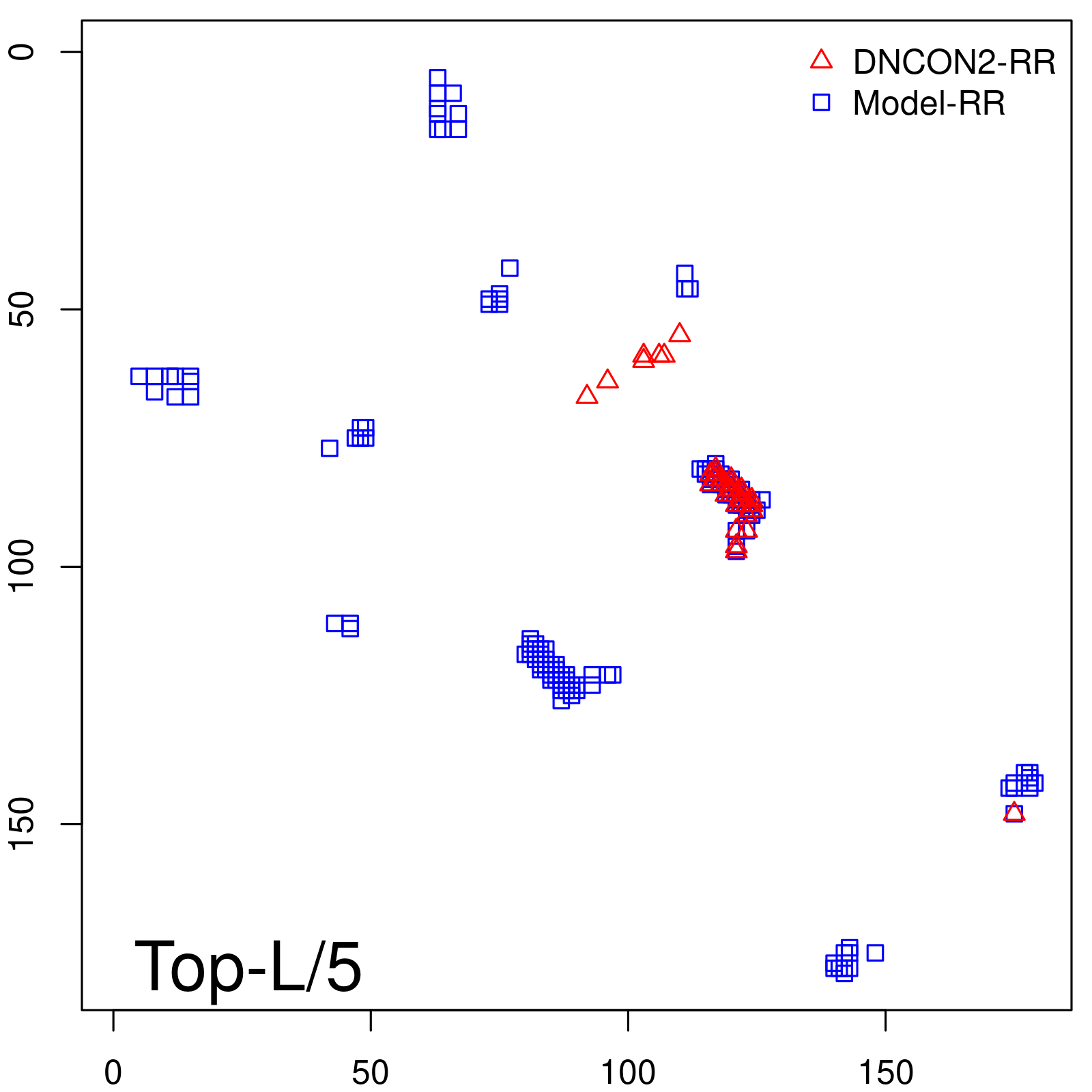
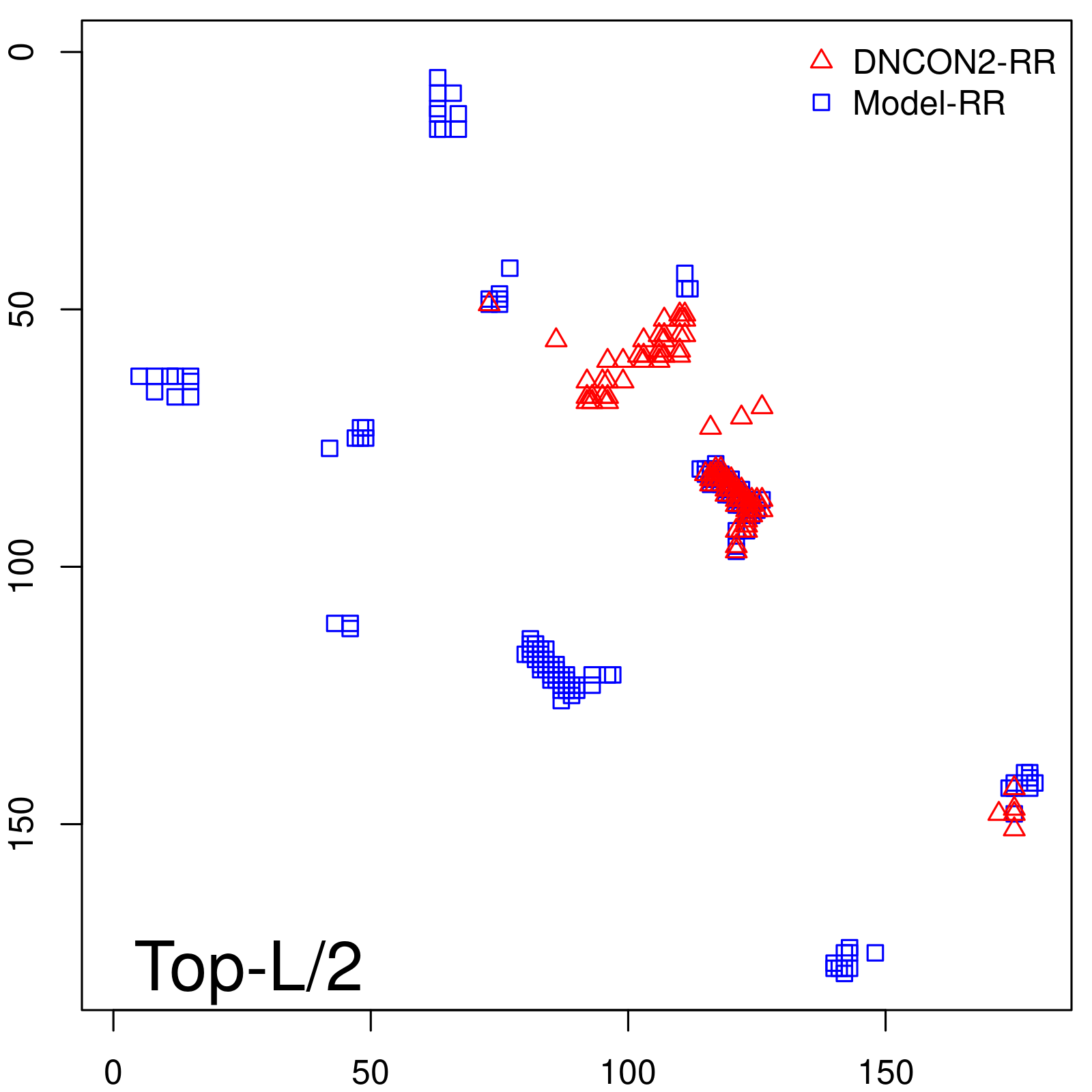
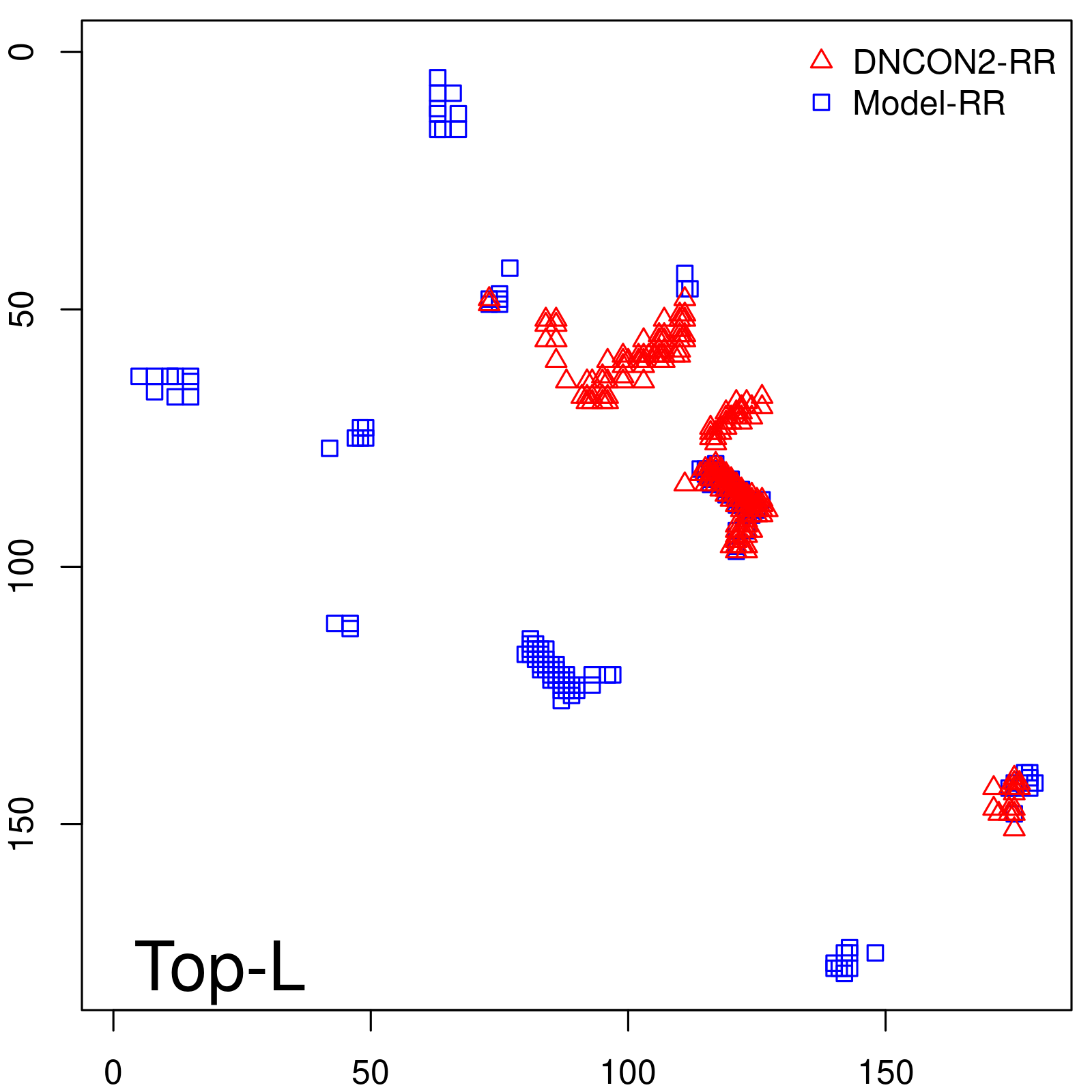
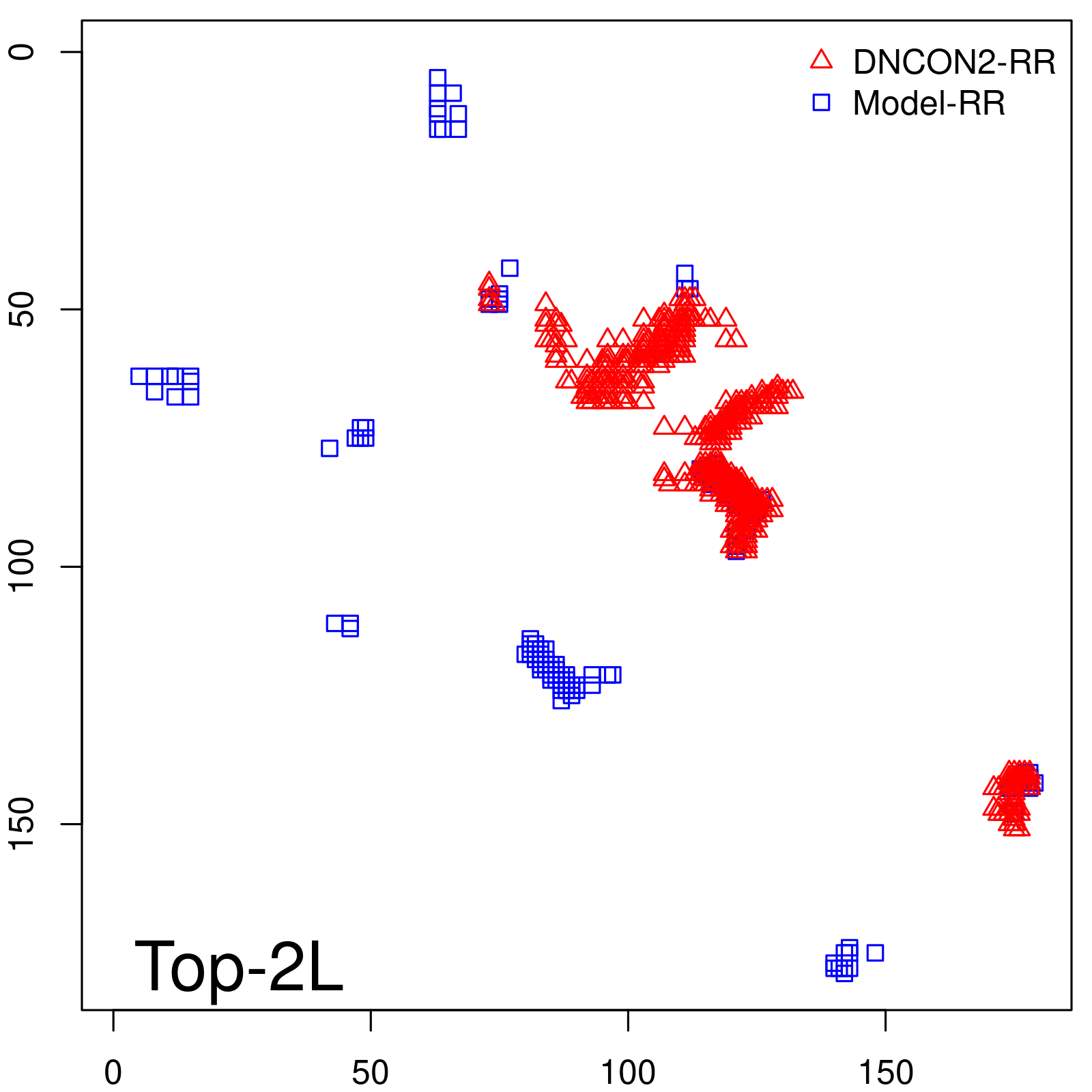
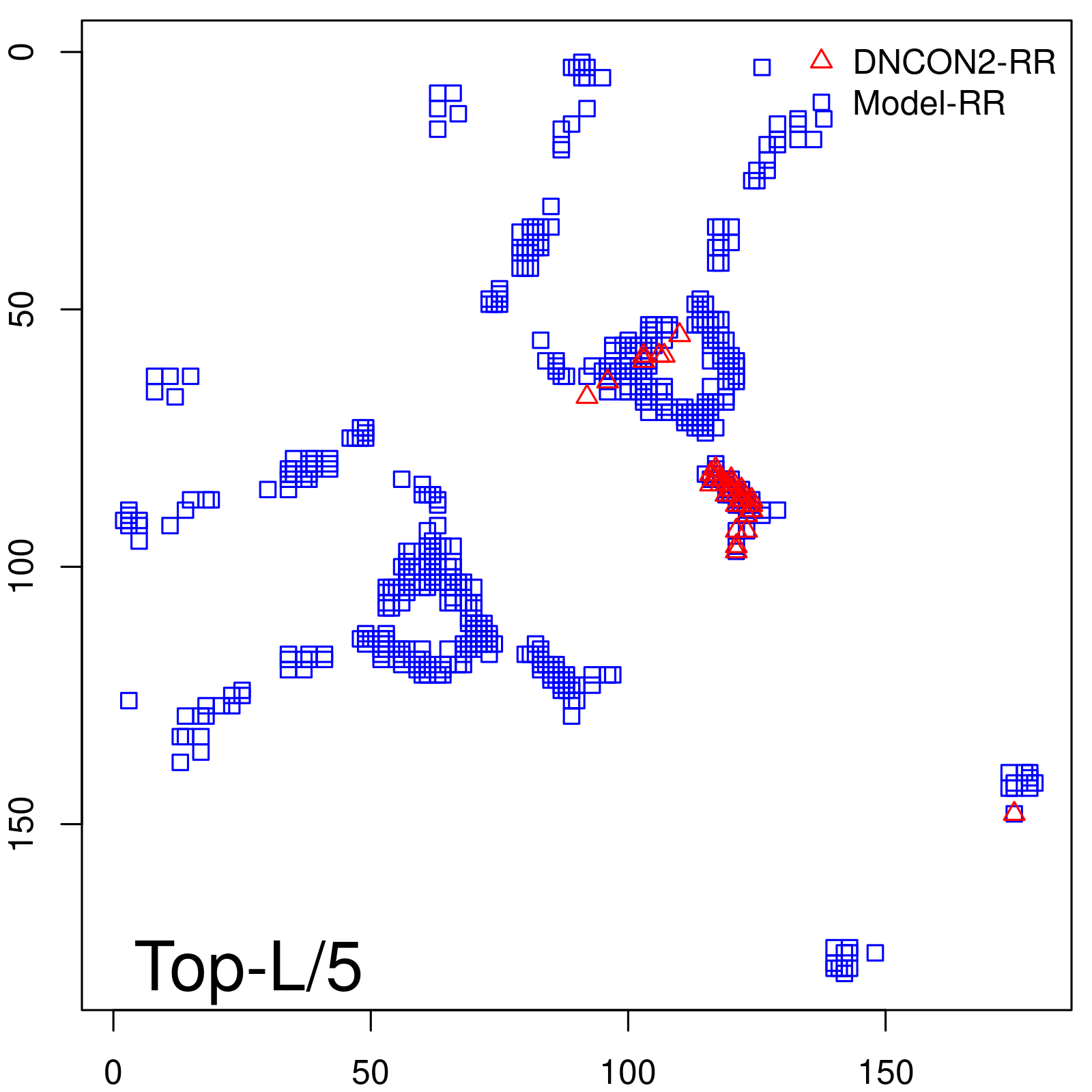
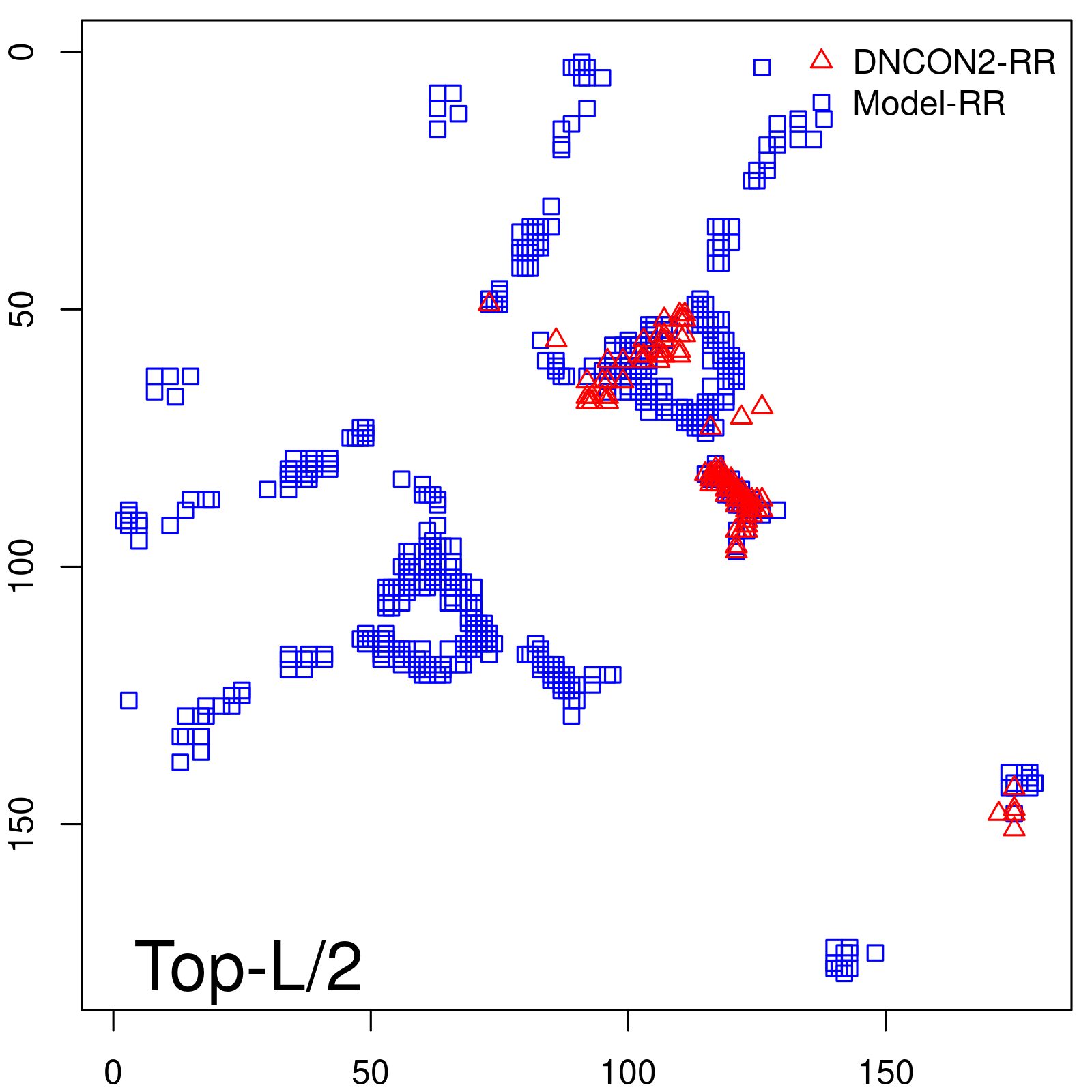
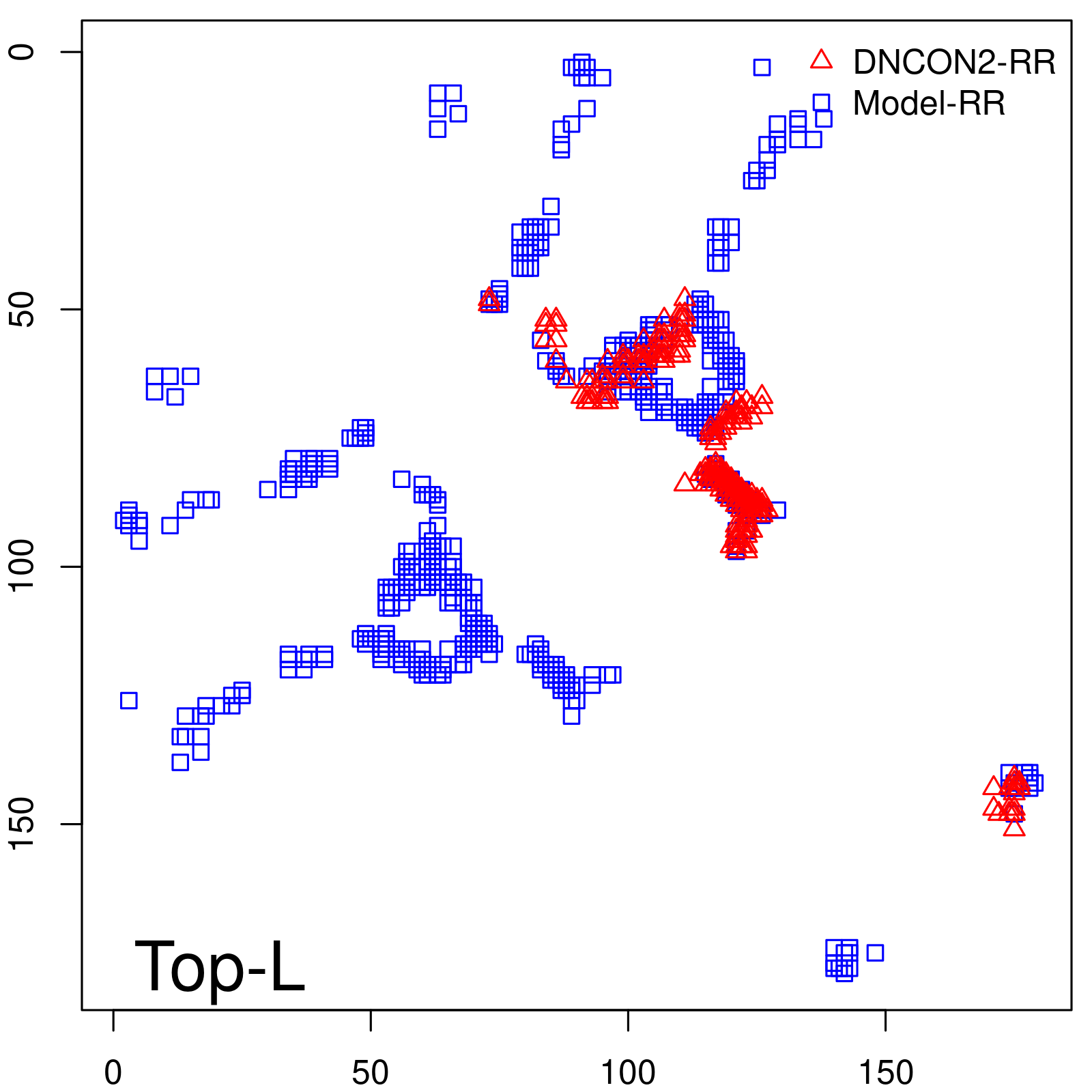
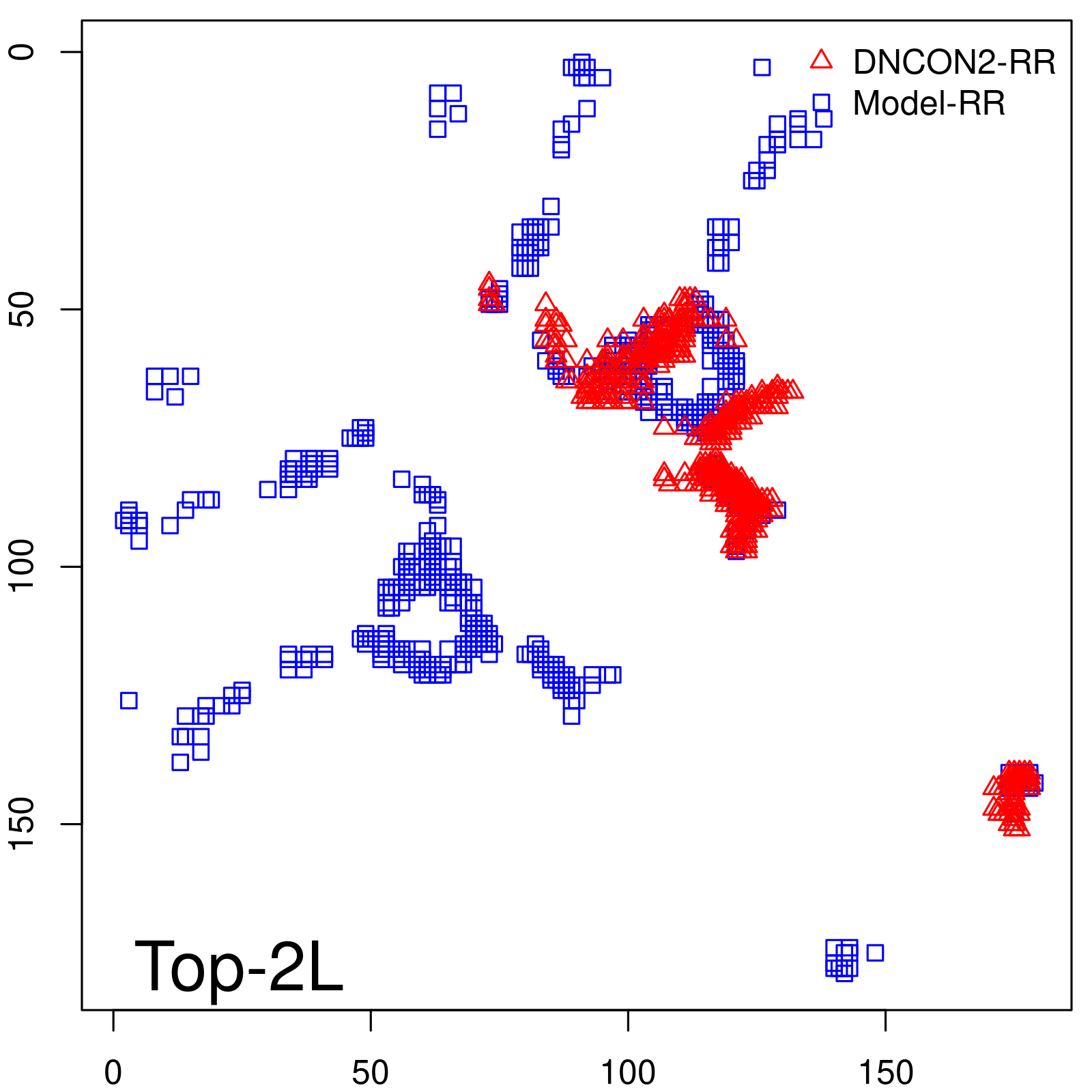
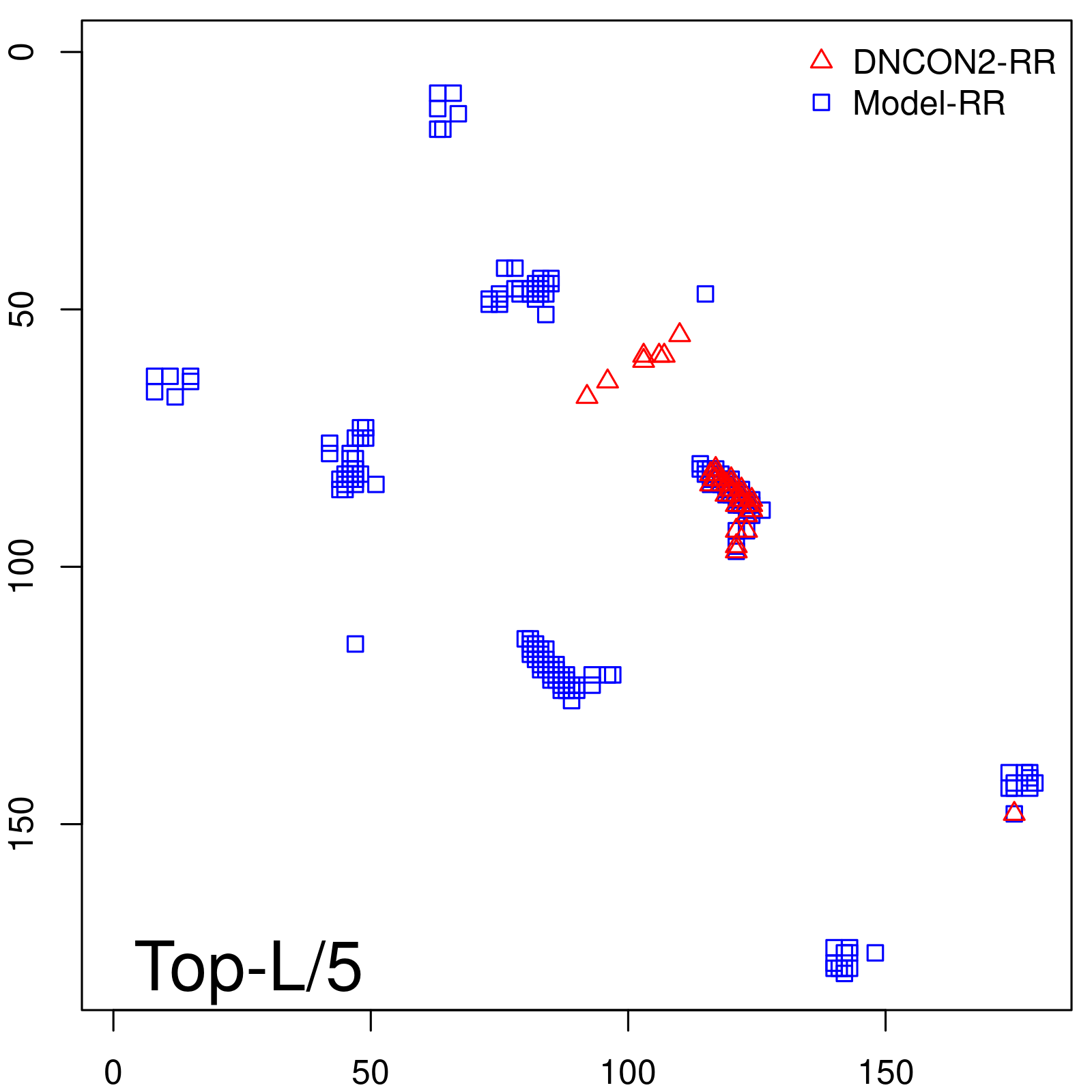
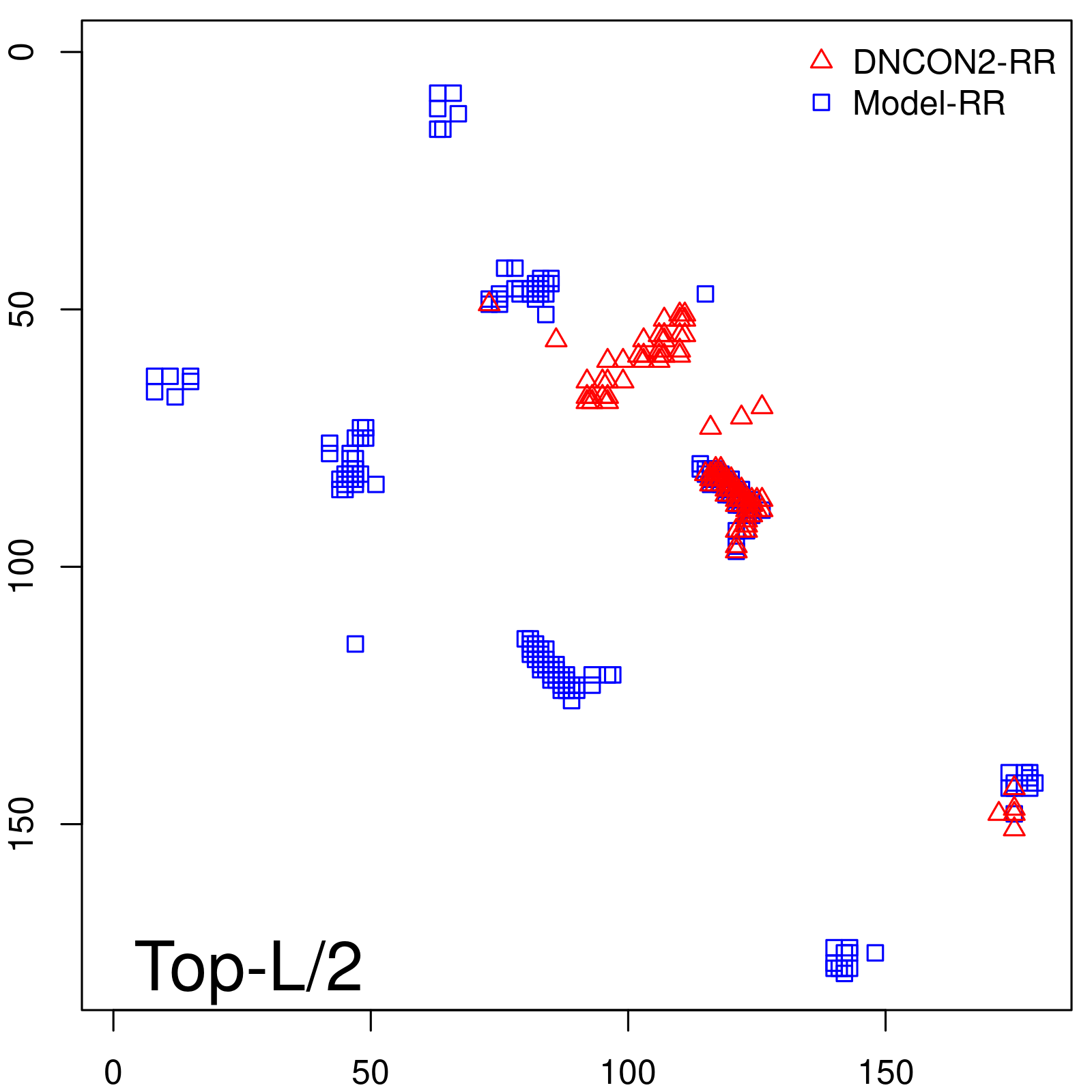
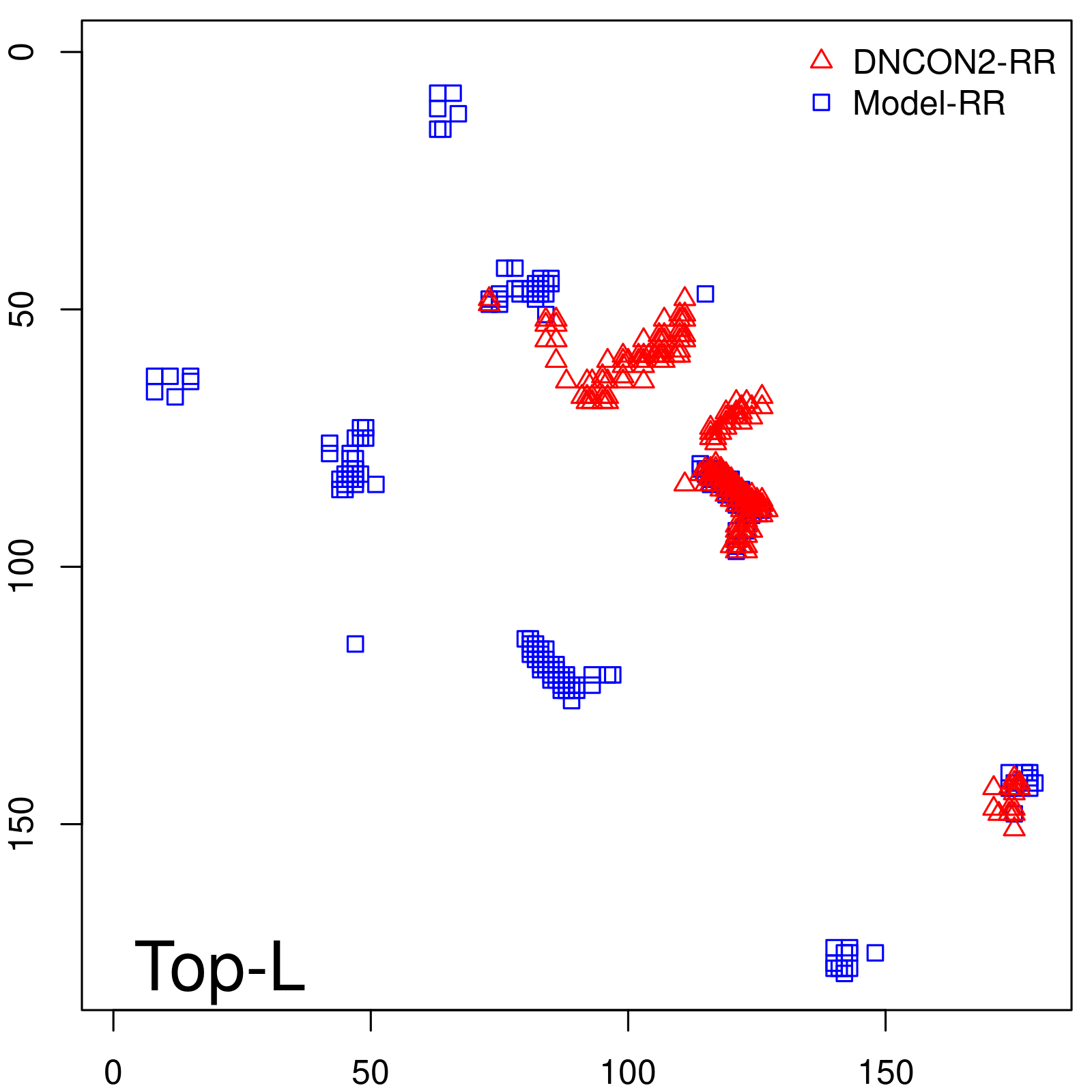
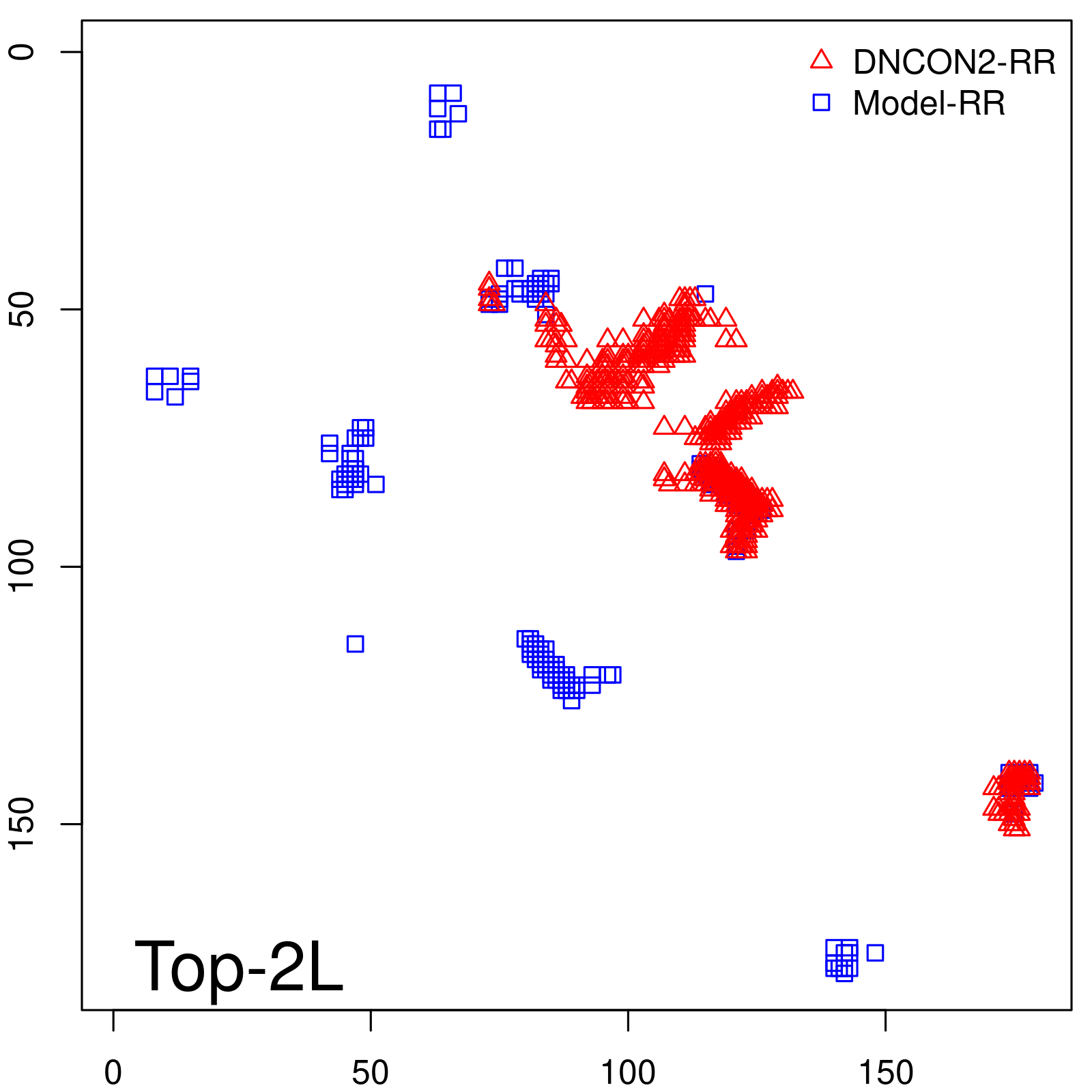
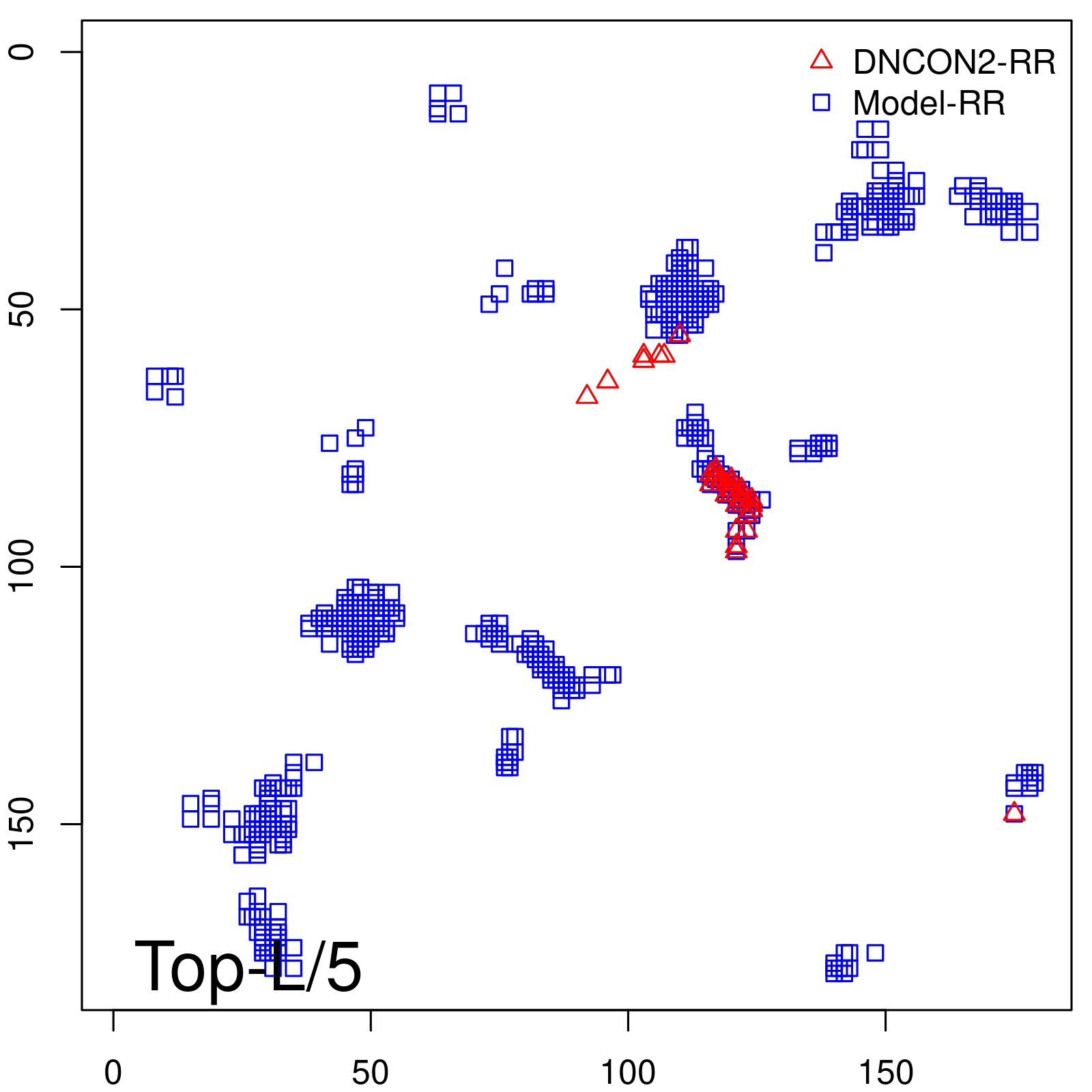
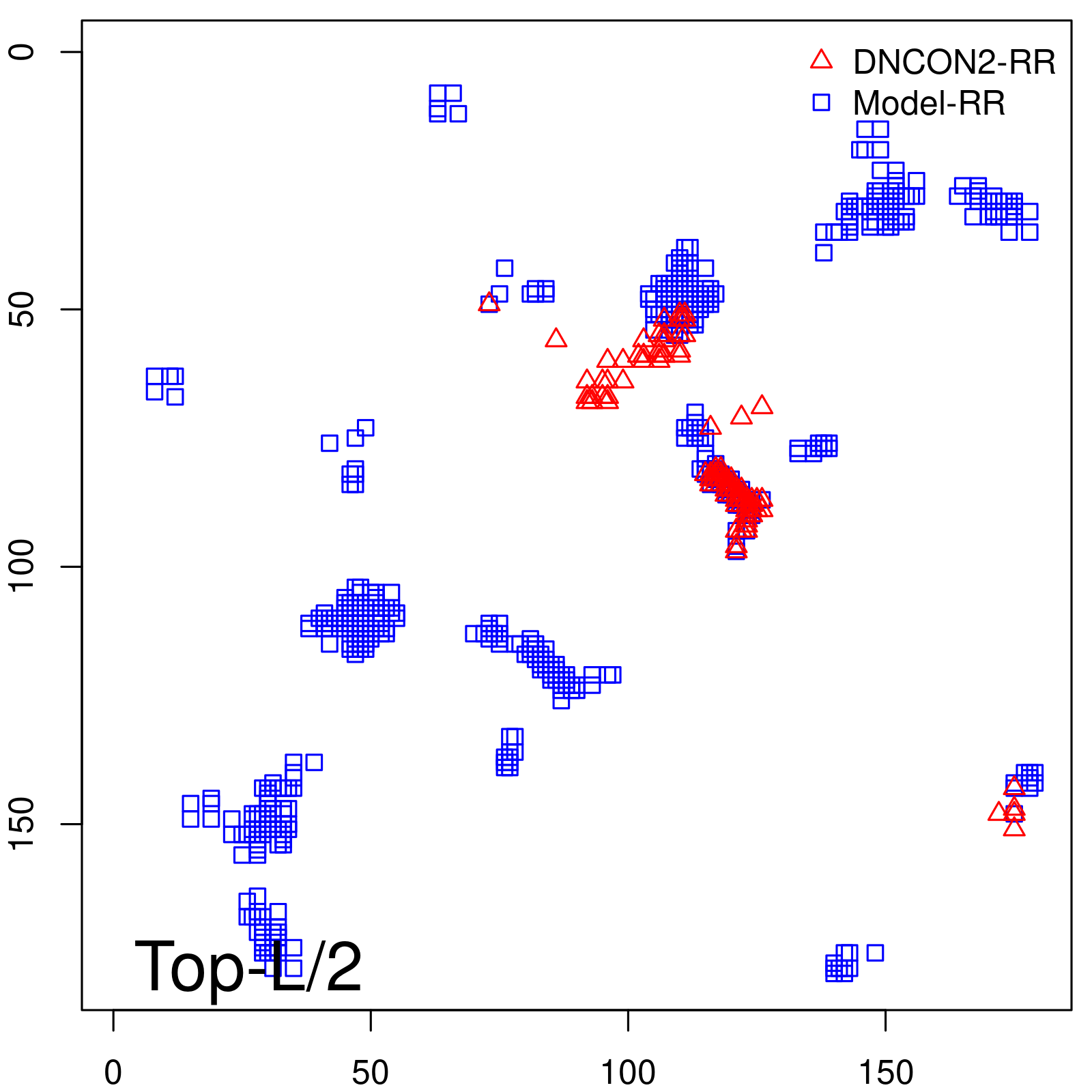
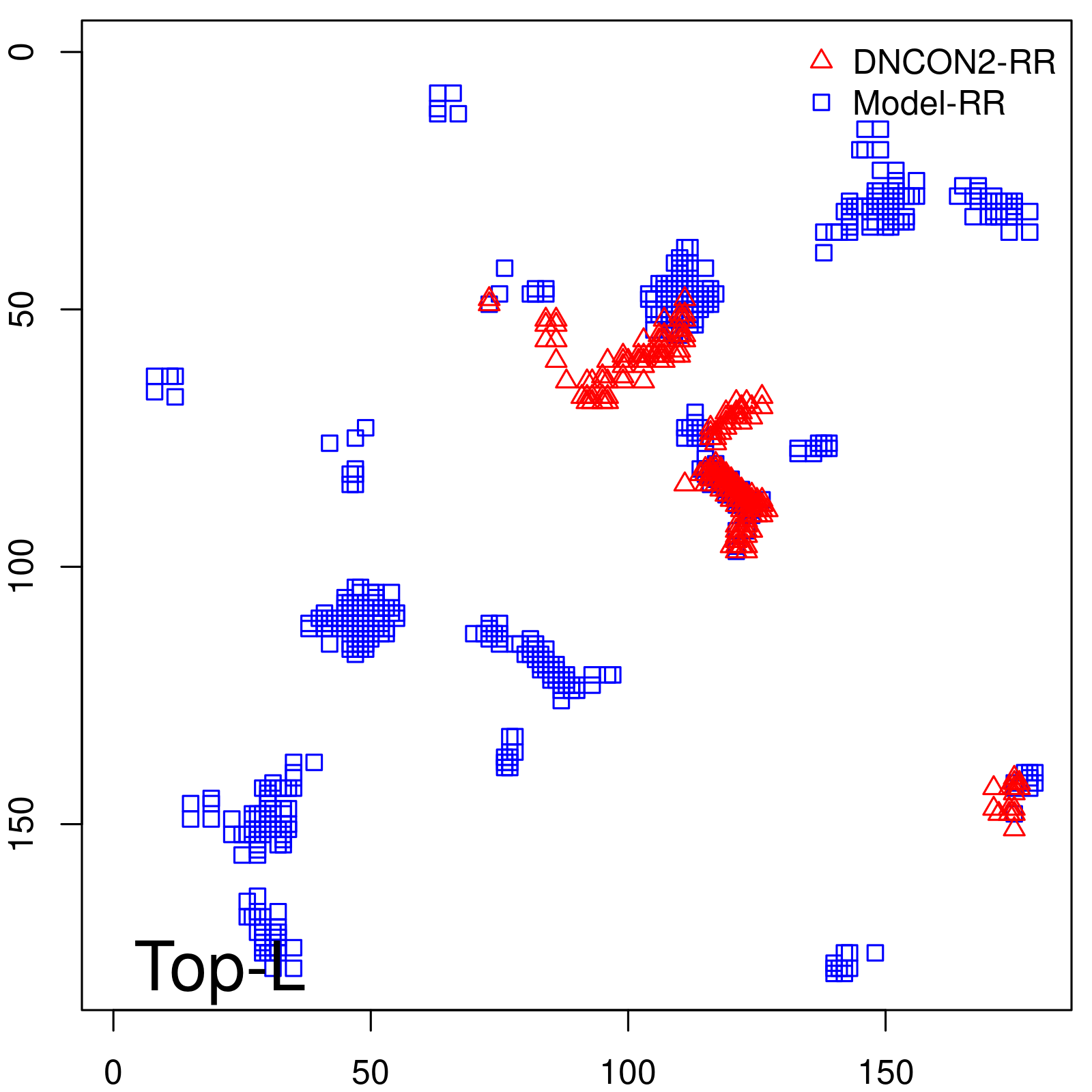
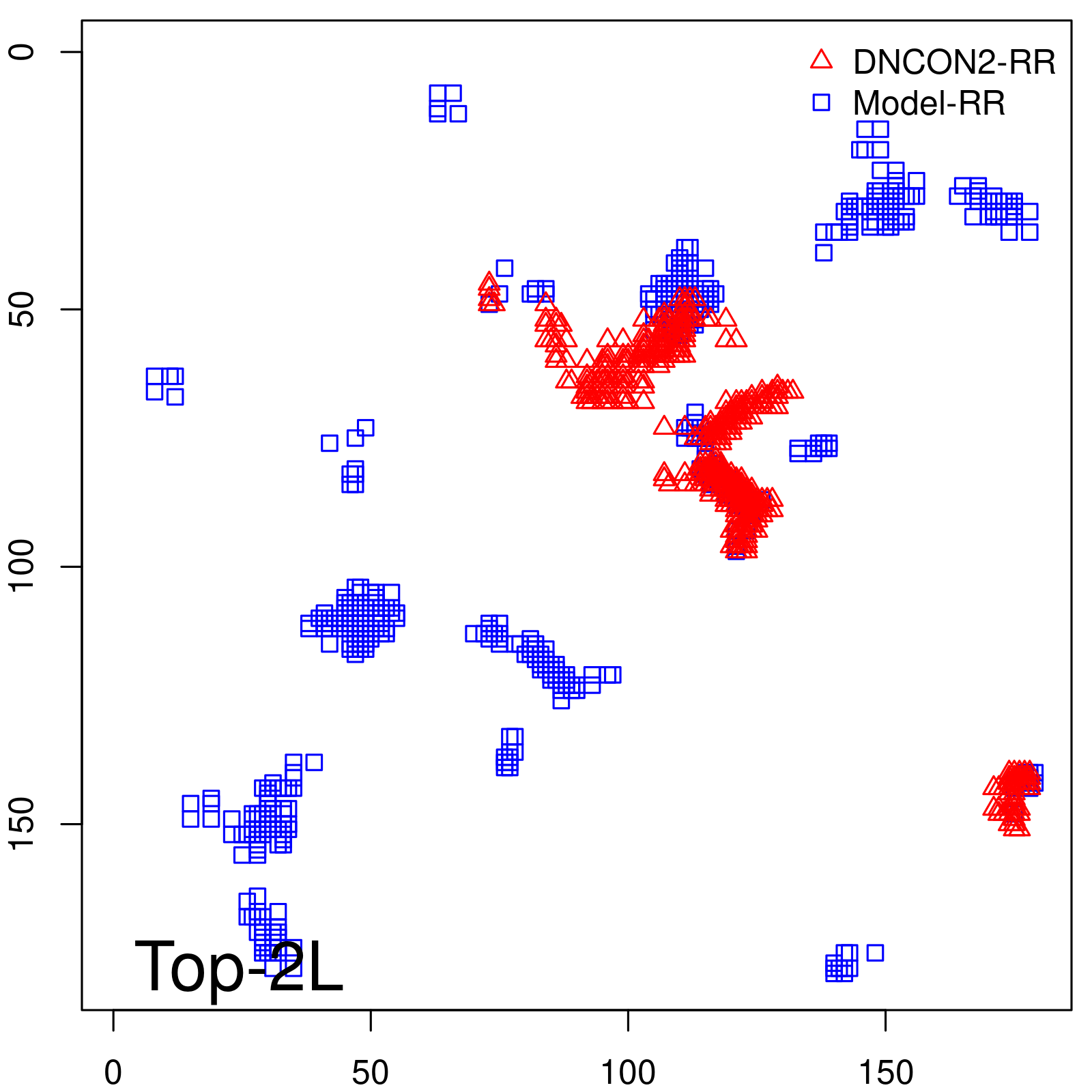
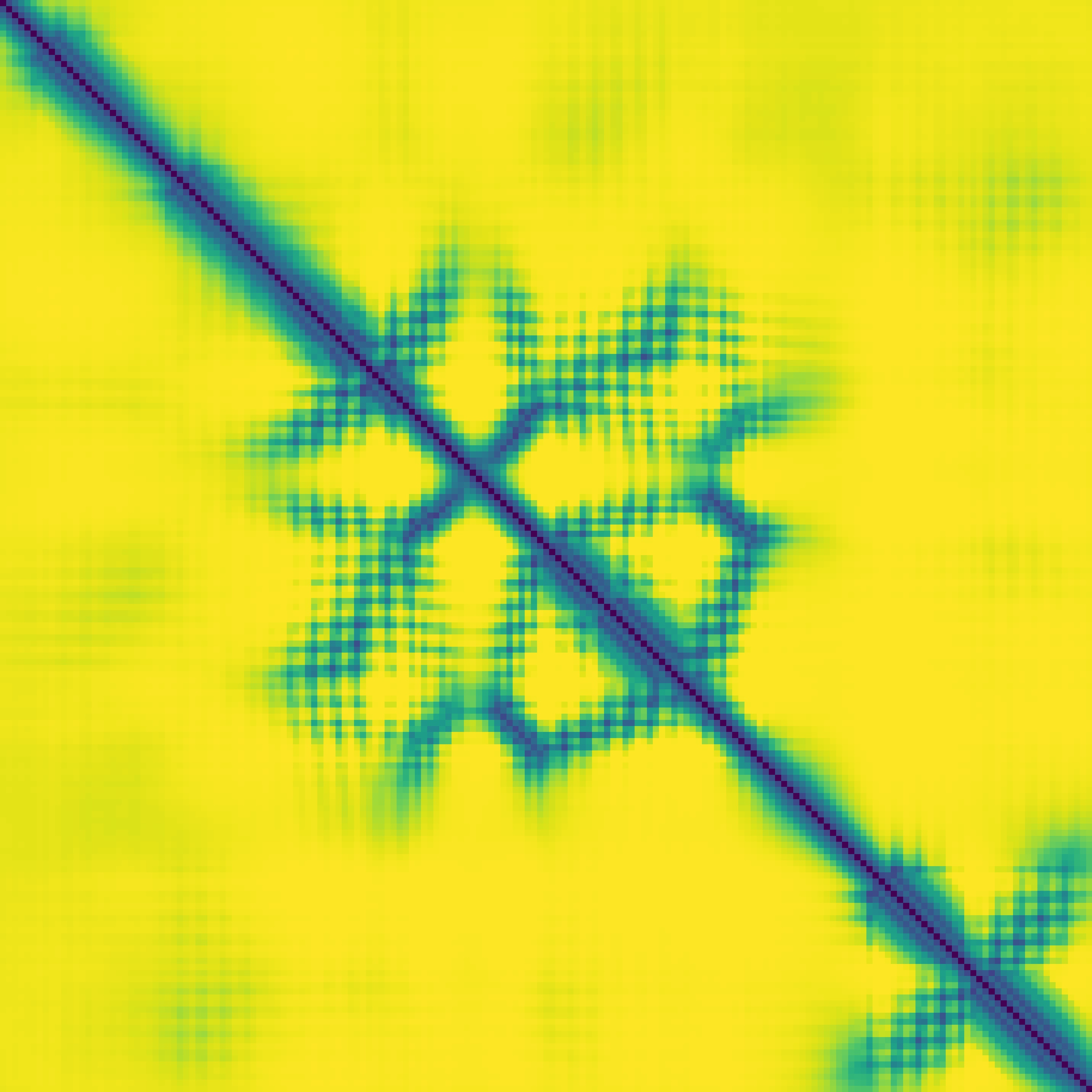
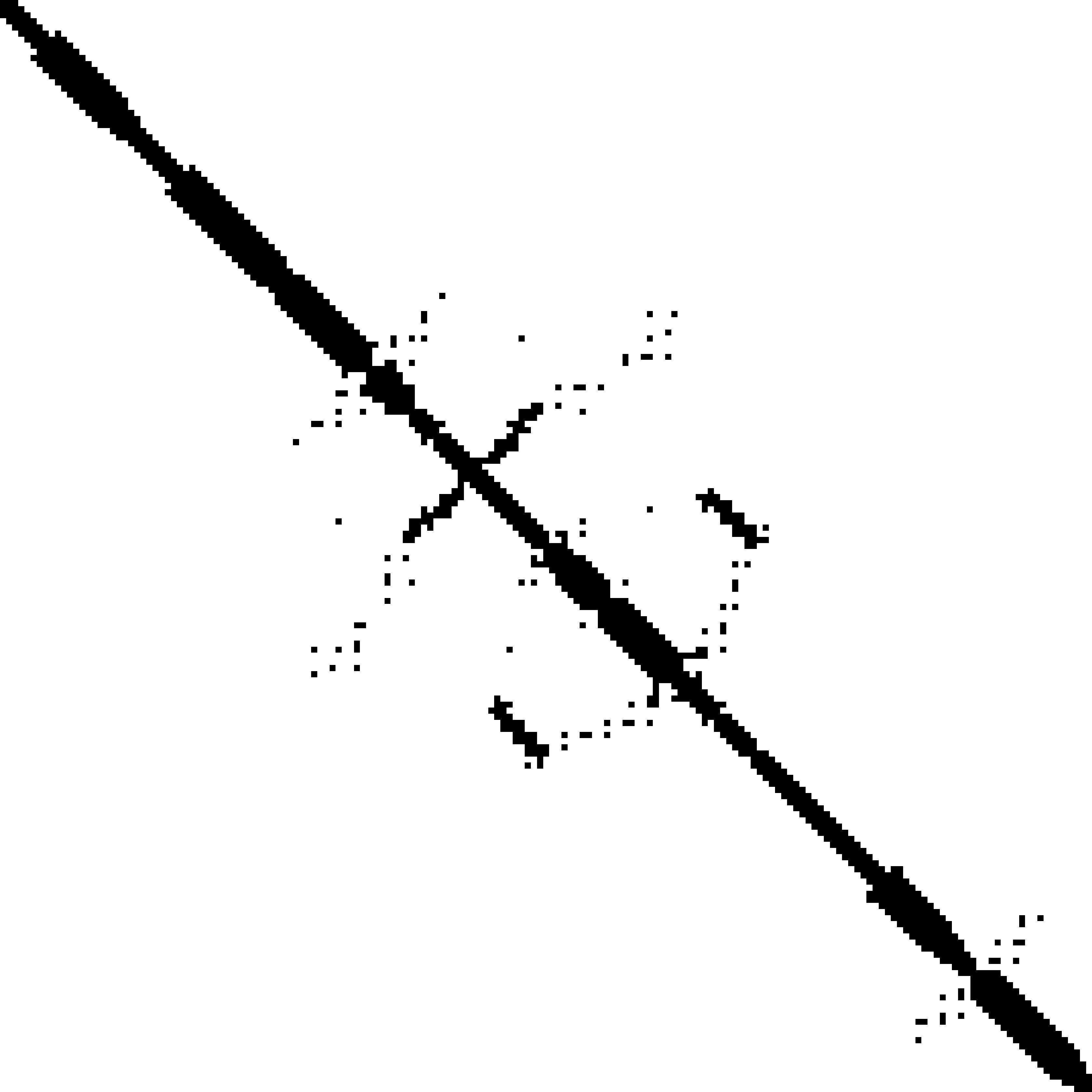
Predicted contact map and distance map
|
Predicted distance map 
|
|
Predicted contact map 
|
Probability to Precision
| Long-Range |
Average probability |
Predicted precision |
| TopL/5 |
0.89 |
88.94 |
| TopL/2 |
0.63 |
66.48 |
Note: The linear model (Average probability vs Predicted precision) was trained on CASP14 dataset
|
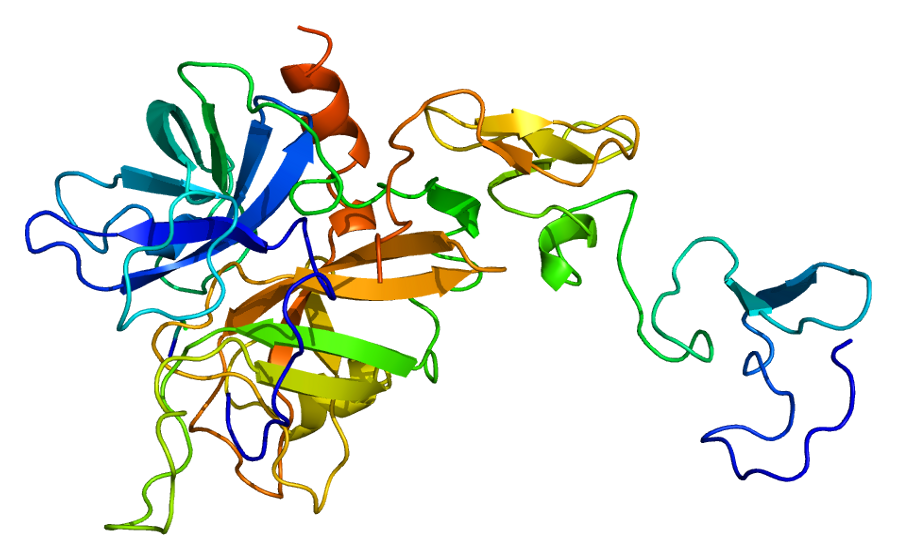
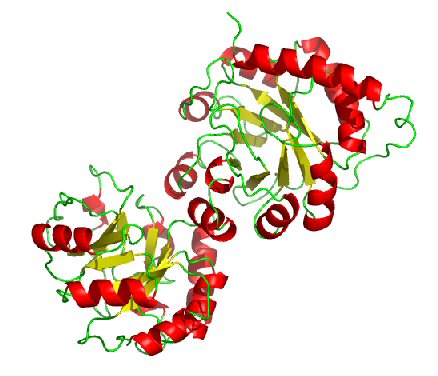
Predicted Top 1 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
80.56 |
| TopL/2 |
44.44 |
| TopL |
25.14 |
| Top2L |
14.53 |
| Alignment |
Number |
| N |
8795 |
| Neff |
1666 |
|
Model comparsion
| Model |
TM-score |
| Model 2 |
0.4109 |
| Model 3 |
0.3752 |
| Model 4 |
0.3863 |
| Model 5 |
0.3649 |
| Average |
0.38433 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 2ztg |
0.46570 |
| 5amr |
0.46510 |
| 2qiz |
0.46113 |
| 6bye |
0.45909 |
| 1ulv |
0.45796 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 2 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
80.56 |
| TopL/2 |
47.78 |
| TopL |
27.37 |
| Top2L |
15.36 |
| Alignment |
Number |
| N |
8795 |
| Neff |
1666 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.4109 |
| Model 3 |
0.3735 |
| Model 4 |
0.3903 |
| Model 5 |
0.3845 |
| Average |
0.38980 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 6bv4 |
0.49832 |
| 6vz9 |
0.48926 |
| 6ufp |
0.48730 |
| 4e36 |
0.48293 |
| 5lg6 |
0.48276 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 3 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
75.00 |
| TopL/2 |
43.33 |
| TopL |
27.37 |
| Top2L |
19.83 |
| Alignment |
Number |
| N |
8795 |
| Neff |
1666 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.3752 |
| Model 2 |
0.3735 |
| Model 4 |
0.3820 |
| Model 5 |
0.3959 |
| Average |
0.38165 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 2z1q |
0.48289 |
| 4ff1 |
0.47843 |
| 4ff4 |
0.47824 |
| 3q24 |
0.47769 |
| 4ff2 |
0.47739 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 4 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
80.56 |
| TopL/2 |
46.67 |
| TopL |
26.26 |
| Top2L |
15.36 |
| Alignment |
Number |
| N |
8795 |
| Neff |
1666 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.3863 |
| Model 2 |
0.3903 |
| Model 3 |
0.3820 |
| Model 5 |
0.3930 |
| Average |
0.38790 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 2xiq |
0.49302 |
| 6s6y |
0.48765 |
| 5t5m |
0.48742 |
| 6oxd |
0.48559 |
| 6req |
0.48334 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 5 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
80.56 |
| TopL/2 |
48.89 |
| TopL |
27.37 |
| Top2L |
18.72 |
| Alignment |
Number |
| N |
8795 |
| Neff |
1666 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.3649 |
| Model 2 |
0.3845 |
| Model 3 |
0.3959 |
| Model 4 |
0.3930 |
| Average |
0.38458 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 4ub9 |
0.45479 |
| 4wgx |
0.45089 |
| 6vkz |
0.44687 |
| 6vl1 |
0.44571 |
| 5cnv |
0.44532 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
References
[1] Hou, J., Wu, T., Cao, R., & Cheng, J. (2019). Protein tertiary structure modeling driven by deep learning and contact distance prediction in CASP13. bioRxiv, 552422.
[2] Li, J., Deng, X., Eickholt, J., & Cheng, J. (2013). Designing and benchmarking the MULTICOM protein structure prediction system. BMC structural biology, 13(1), 2.
[3] Cheng, J., Li, J., Wang, Z., Eickholt, J., & Deng, X. (2012). The MULTICOM toolbox for protein structure prediction. BMC bioinformatics, 13(1), 65.
[4] Wang, Z., Eickholt, J., & Cheng, J. (2010). MULTICOM: a multi-level combination approach to protein structure prediction and its assessments in CASP8. Bioinformatics, 26(7), 882-888.
 )
) )
)
 )
) )
) )
) )
) )
)