P9WM57
multicom
P9WM57
full_length
P9WM57
Results of Structure Prediction for Target Name: P9WM57 ( Click  )
)
Domain Boundary prediction ( View  )
)

>P9WM57: 1-451
| 1-60: |
M | C | S | T | R | E | E | I | T | E | A | F | A | S | L | A | T | A | L | S | R | V | L | G | L | T | F | D | A | L | T | T | P | E | R | L | A | L | L | E | H | C | E | T | A | R | R | Q | L | P | S | V | E | H | T | L | I | N | Q | I |
| 61-119: |
G | E | Q | S | T | E | E | E | L | G | G | K | L | G | L | T | L | A | D | R | L | R | I | T | R | S | E | A | K | R | R | V | A | E | A | A | D | L | G | Q | R | R | A | L | T | G | E | P | L | P | P | L | L | T | A | T | A | K | A | Q |
| 121-179: |
R | H | G | L | I | G | D | G | H | V | E | V | I | R | A | F | V | H | R | L | P | S | W | V | D | L | K | T | L | E | K | A | E | R | D | L | A | K | Q | A | T | Q | Y | R | P | D | Q | L | A | K | L | A | A | R | I | M | D | C | L | N |
| 181-239: |
P | D | G | D | Y | T | D | E | D | R | A | R | R | R | G | L | T | L | G | K | Q | D | V | D | G | M | S | R | L | S | G | Y | V | T | P | E | L | R | A | T | I | E | A | V | W | A | K | L | A | A | P | G | M | C | N | P | E | Q | K | A |
| 241-299: |
P | C | V | N | G | A | P | S | K | E | Q | A | R | R | D | T | R | S | C | P | Q | R | N | H | D | A | L | N | A | E | L | R | S | L | L | T | S | G | N | L | G | Q | H | N | G | L | P | A | S | I | I | V | T | T | T | L | K | D | L | E |
| 301-359: |
A | A | A | G | A | G | L | T | G | G | G | T | I | L | P | I | S | D | V | I | R | L | A | R | H | A | N | H | Y | L | A | I | F | D | R | G | K | A | L | A | L | Y | H | T | K | R | L | A | S | P | A | Q | R | I | M | L | Y | A | K | D |
| 361-419: |
S | G | C | S | A | P | G | C | D | V | P | G | Y | Y | C | E | V | H | H | V | T | P | Y | A | Q | C | R | N | T | D | V | N | D | L | T | L | G | C | G | G | H | H | P | L | A | E | R | G | W | T | T | R | K | N | A | H | G | D | T | E |
| 421-451: |
W | L | P | P | P | H | L | D | H | G | Q | P | R | V | N | T | F | H | H | P | E | K | L | L | A | D | D | E | G | D | P |
| 1-60: |
C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | H | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H |
| 61-119: |
H | H | H | C | C | C | H | H | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H |
| 121-179: |
H | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C | H | H | H | H | H | H | H | H | H | H | H | H | H | H | C | C |
| 181-239: |
C | C | C | C | C | C | C | H | H | H | H | H | C | C | C | E | E | E | E | C | C | C | C | C | C | C | E | E | E | E | C | C | C | C | H | H | H | H | H | H | H | H | H | H | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C |
| 241-299: |
C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | H | H | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | E | E | E | E | E | C | C | C | C | C |
| 301-359: |
C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C | C | C | C | C | E | C | C | C | C | C | C | C | C | E | E | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C |
| 361-419: |
C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | C | H | H | H | H | C | C | C | C | C | C | C | C | C | C | C | E | E | E | E | E | C | C | C | C | C | E | E |
| 421-451: |
E | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | C | H | H | C | C | C | C | C | C | C | C |
|
| | H(Helix): 183(40.58%) | E(Strand): 24(5.32%) | C(Coil): 244(54.1%) |
| 1-60: |
E | E | B | E | E | E | E | B | E | E | B | B | E | B | B | E | E | B | B | E | E | B | B | E | B | E | B | E | E | B | E | E | E | E | B | B | E | B | B | E | B | B | E | E | B | B | E | B | B | E | B | B | B | B | B | B | B | B | B | B |
| 61-119: |
B | E | E | E | B | B | E | E | B | E | B | E | B | B | E | B | B | B | E | B | B | E | B | B | E | E | E | B | E | B | B | B | E | B | B | E | E | B | B | E | E | E | E | B | E | E | E | E | B | E | E | E | B | E | E | B | B | E | B | B |
| 121-179: |
E | E | E | E | B | E | B | E | B | B | E | B | B | B | E | B | B | B | E | B | B | E | B | B | E | B | E | B | B | E | E | B | B | E | B | B | B | E | B | B | E | E | B | E | B | E | E | B | E | E | B | B | E | E | B | B | E | B | B | B |
| 181-239: |
E | E | E | E | E | E | E | B | E | E | B | B | B | B | B | B | B | B | B | E | E | E | B | E | B | B | B | B | B | B | B | B | B | E | B | E | B | B | B | B | B | B | B | B | B | B | E | B | B | B | E | B | B | B | E | E | B | E | E | B |
| 241-299: |
B | E | B | E | E | E | E | B | E | B | B | B | B | E | E | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | E | B | E | E | B | E | E | E | E | E | B | E | B | B | B | B | B | B | B | B | B | B | B | B | B |
| 301-359: |
E | E | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | B | E | E | E | E | B | B | E | B | B | B | E | E | B | E | B | E | E | E | B | B | B | B | B | B | B | B | B |
| 361-419: |
E | B | B | B | B | B | B | B | E | B | B | B | B | B | B | B | B | B | B | B | B | E | B | B | E | E | B | E | B | B | B | E | B | B | B | B | B | B | B | B | B | B | B | B | E | B | E | E | B | B | B | B | E | E | E | E | B | B | B | B |
| 421-451: |
B | B | B | B | E | E | B | B | E | E | E | E | E | E | B | E | B | B | B | B | E | E | B | B | E | E | E | E | E | E | E |
|
| | e(Exposed): 167(37.03%) | b(Buried): 284(62.97%) |
| 1-60: |
T | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 61-119: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 121-179: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 181-239: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | T | T | T | T | T | T | T |
| 241-299: |
T | T | T | T | T | T | T | T | T | T | T | T | T | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 301-359: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 361-419: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 421-451: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | T | T | T | T | T | T | T |
|
| | N(Normal): 423(93.79%) | T(Disorder): 28(6.21%) |
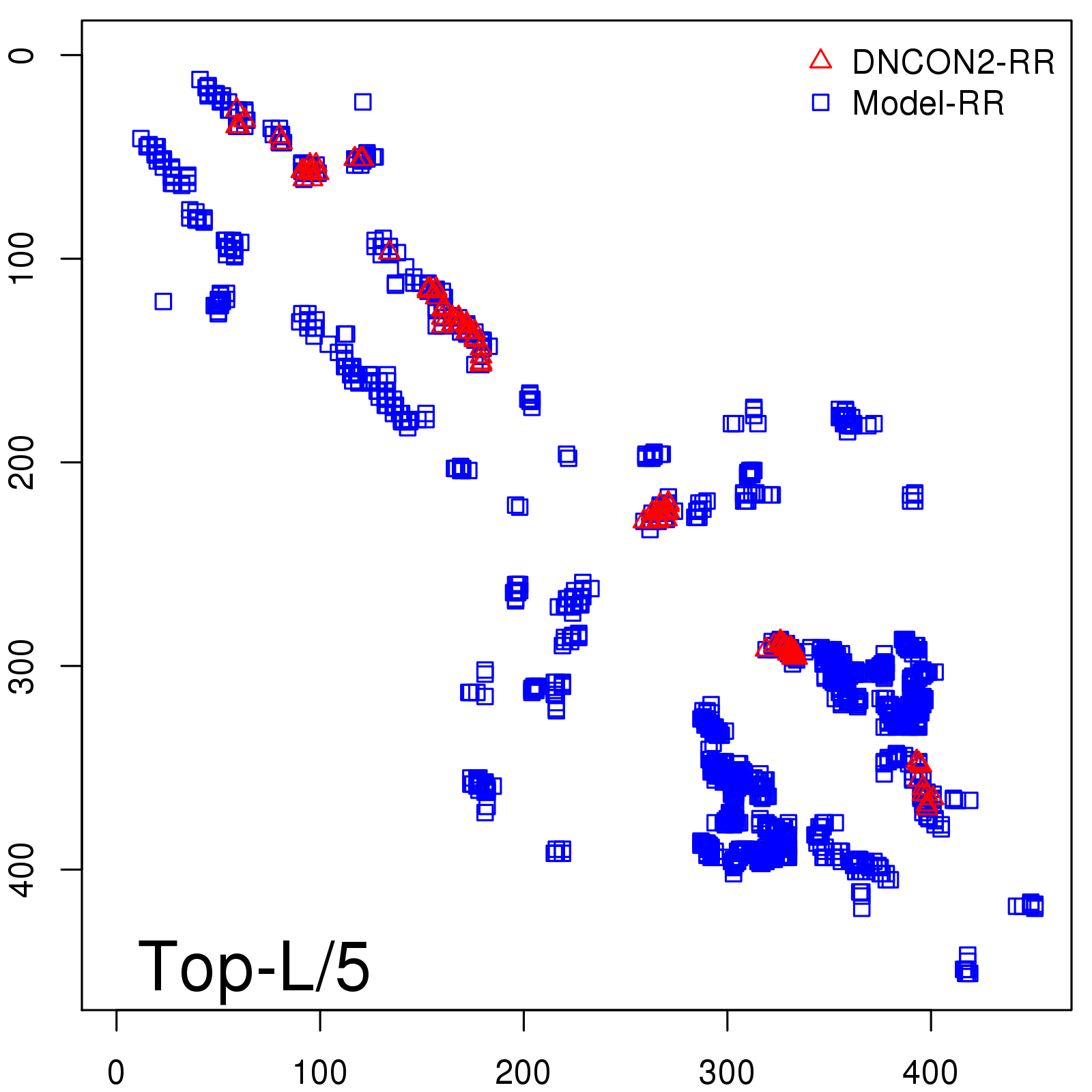
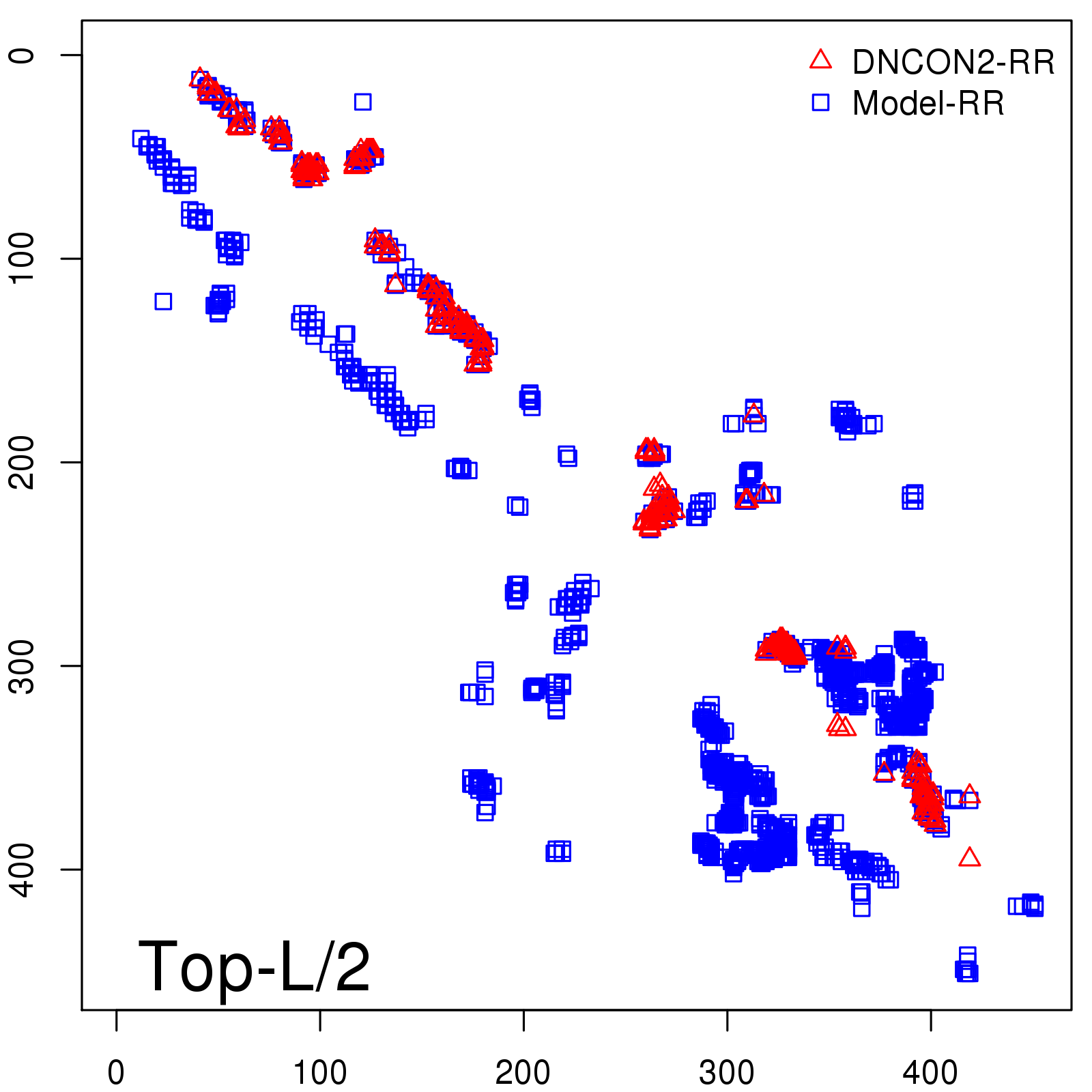
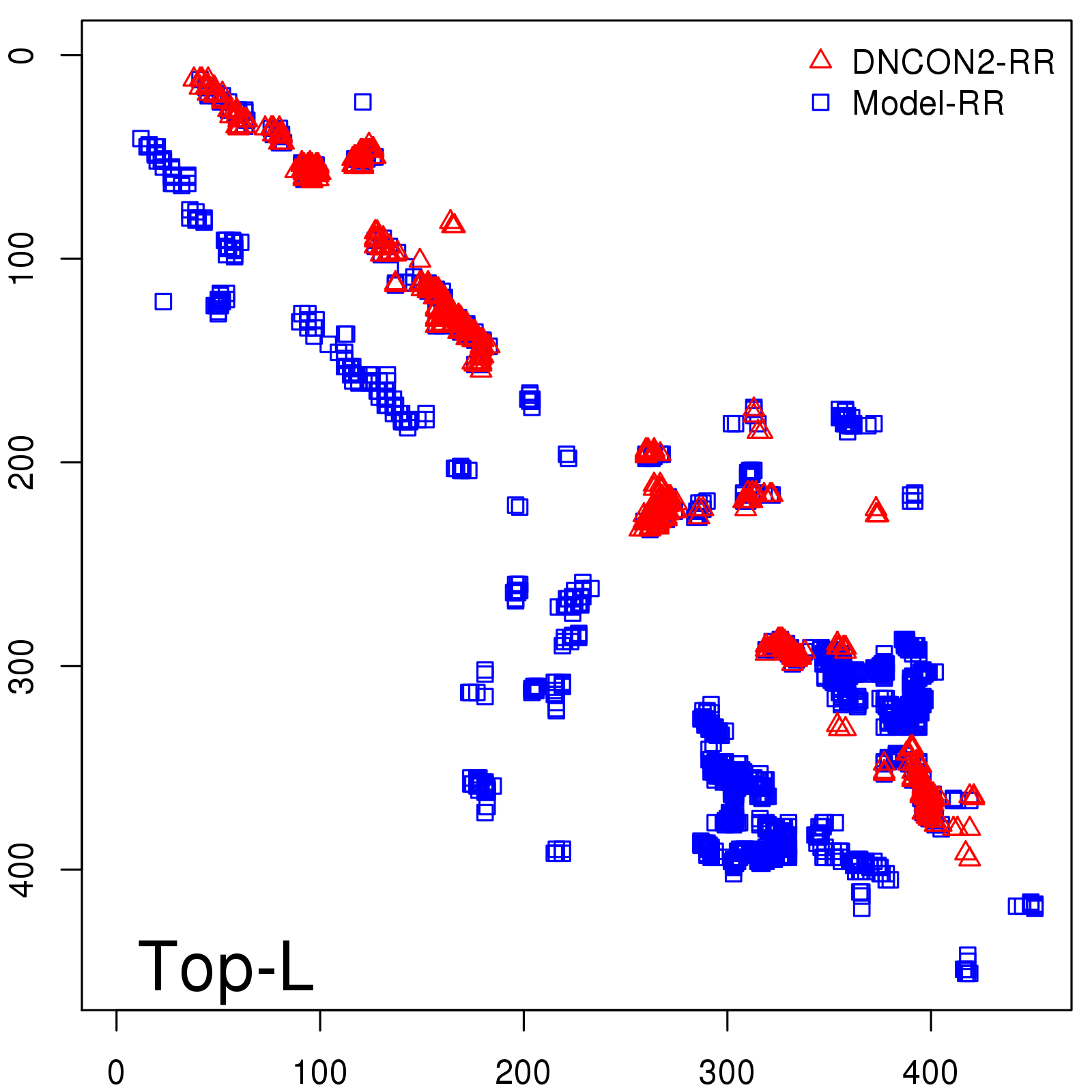
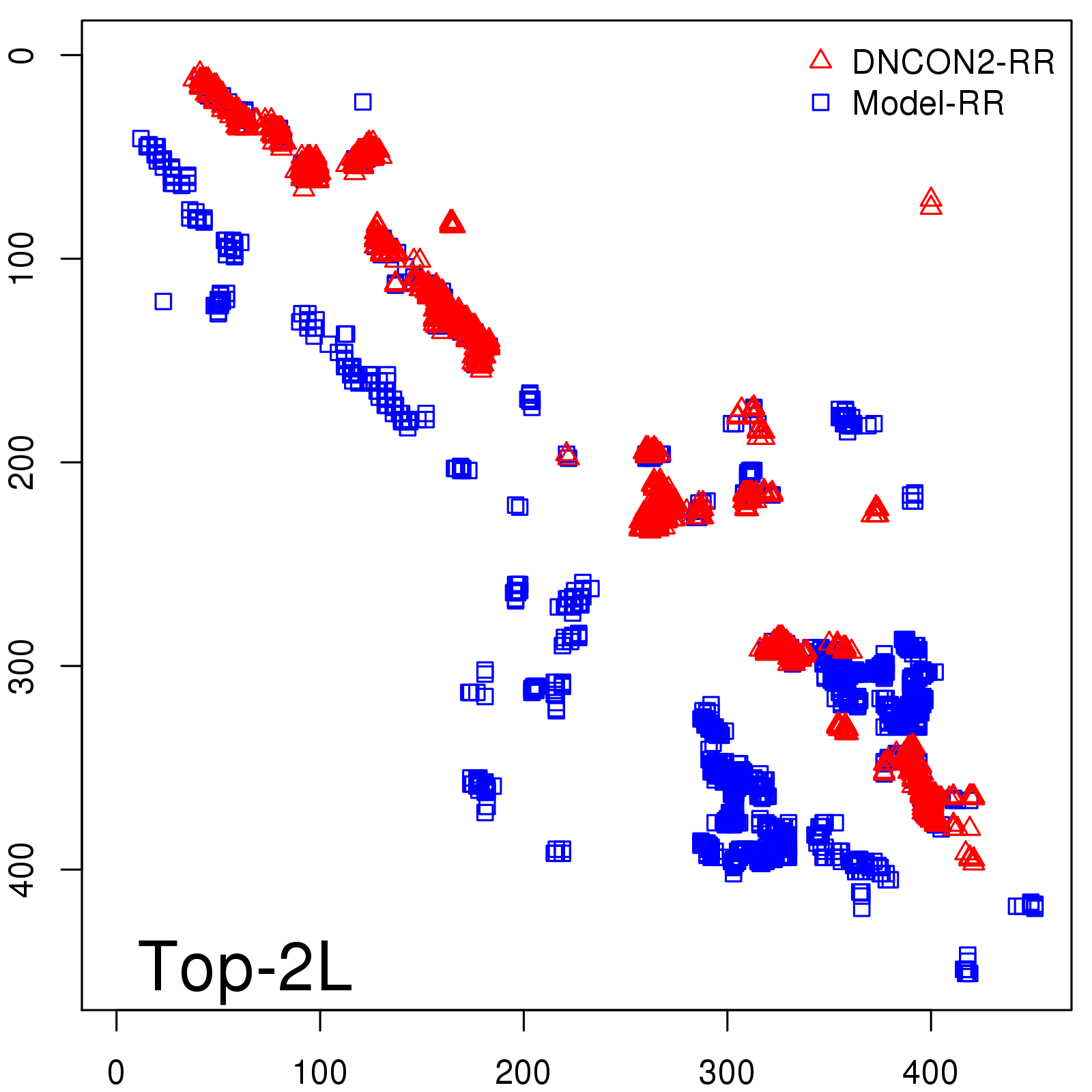
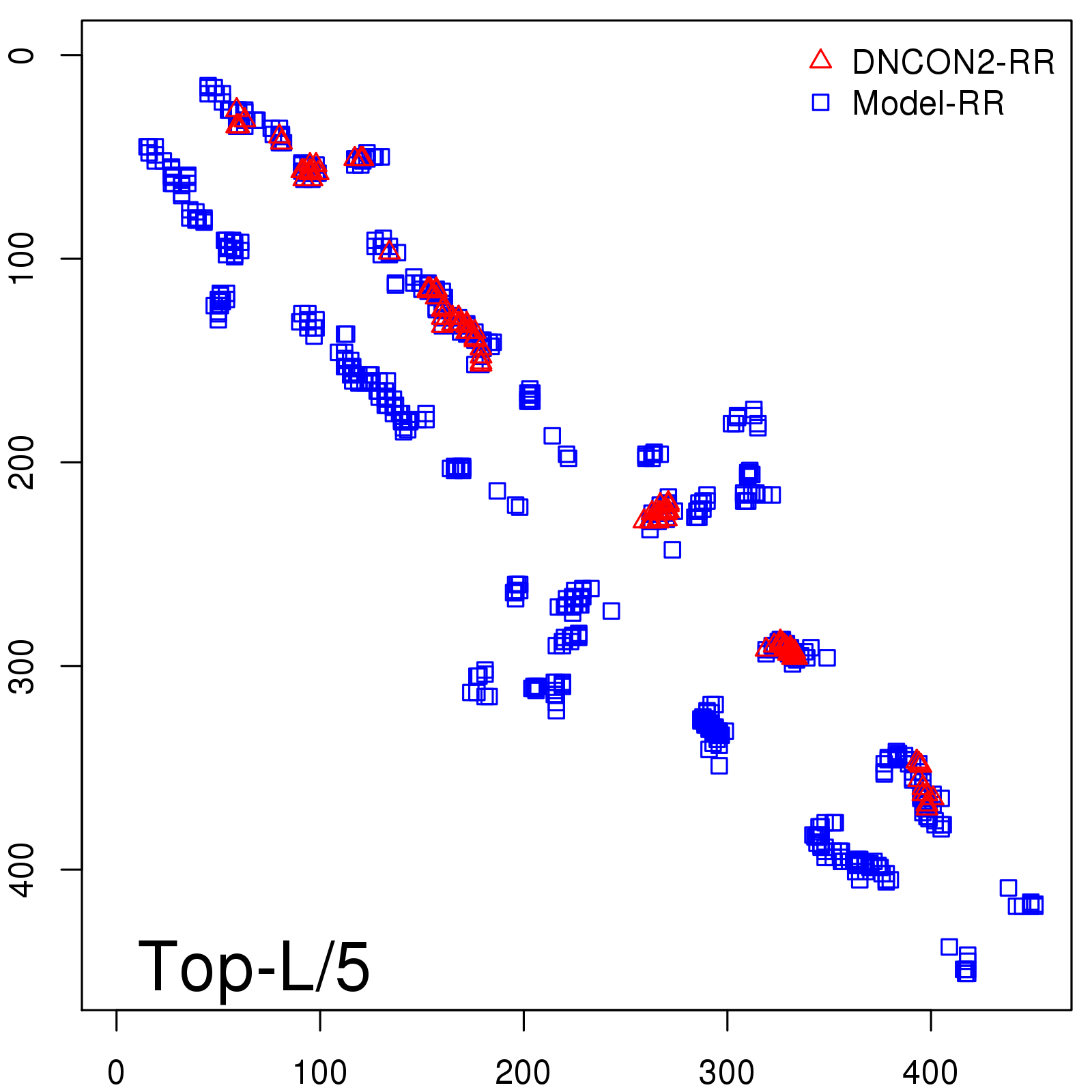
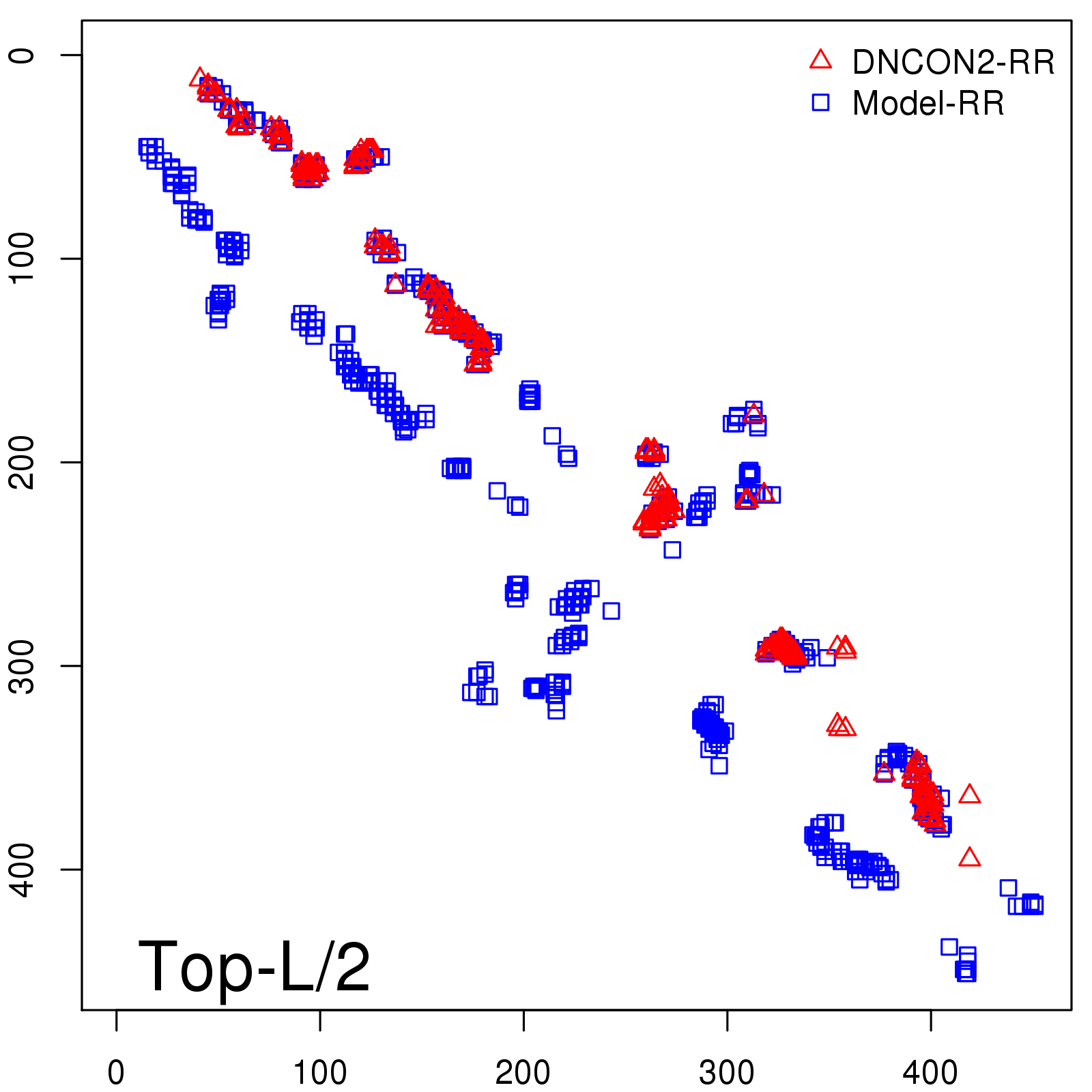
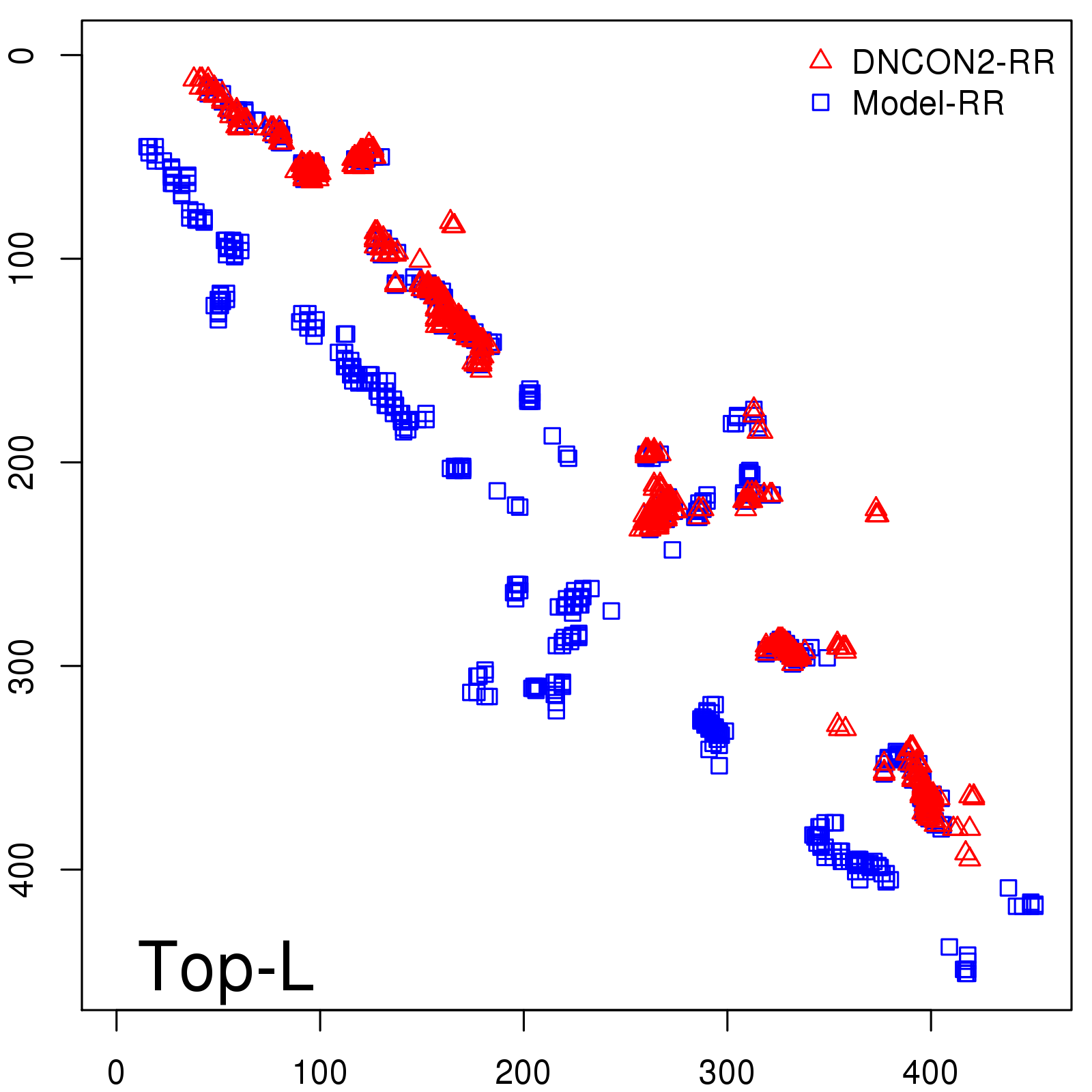
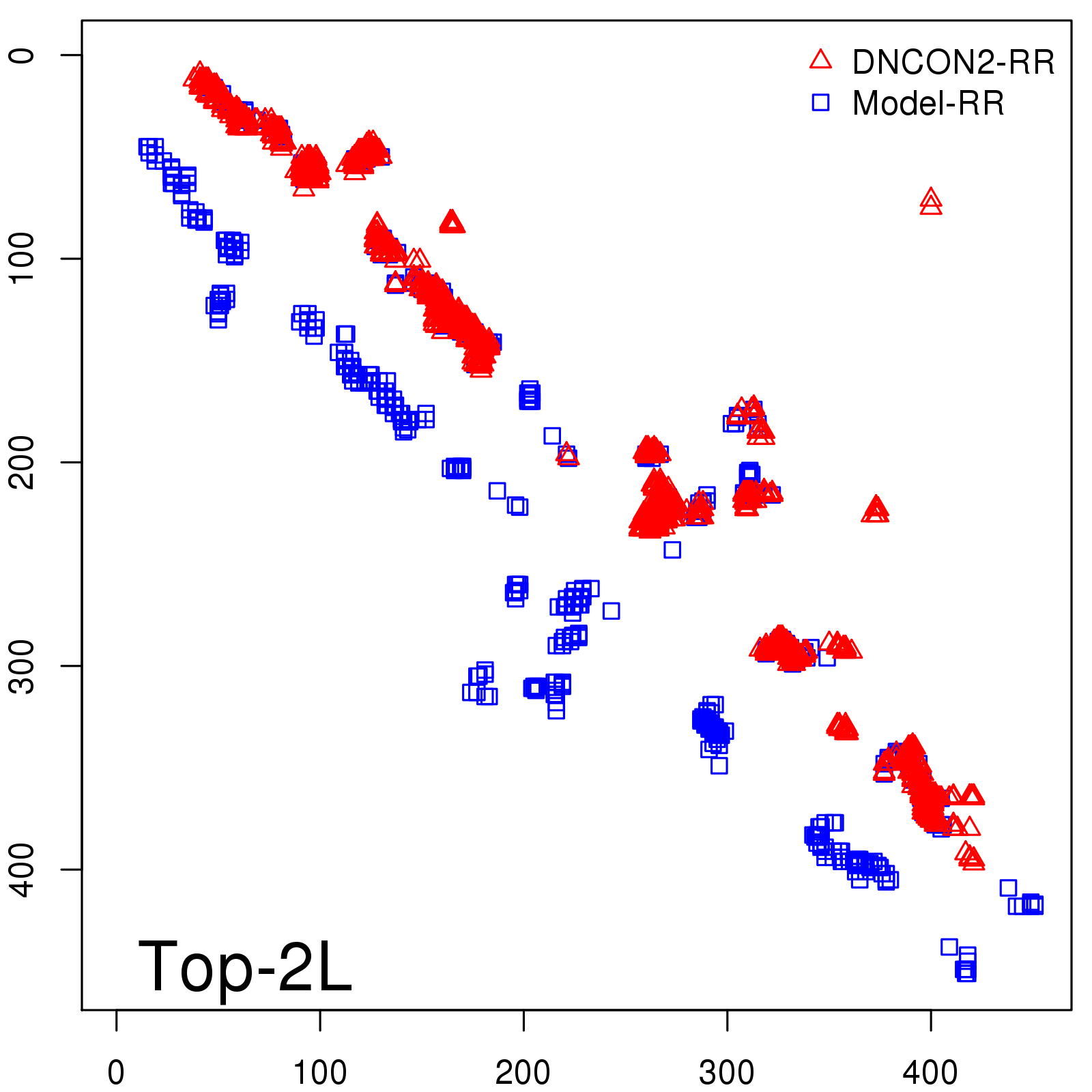
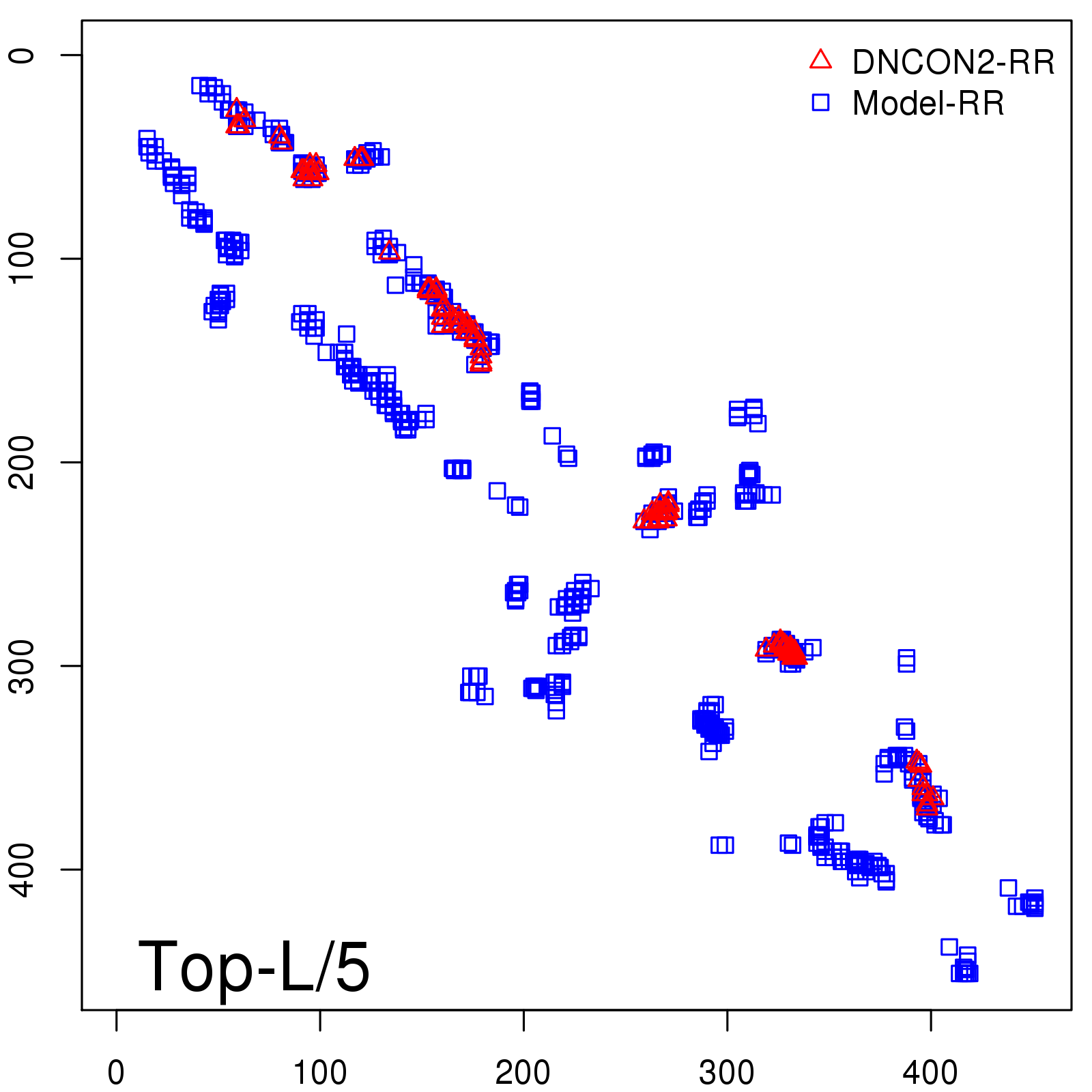
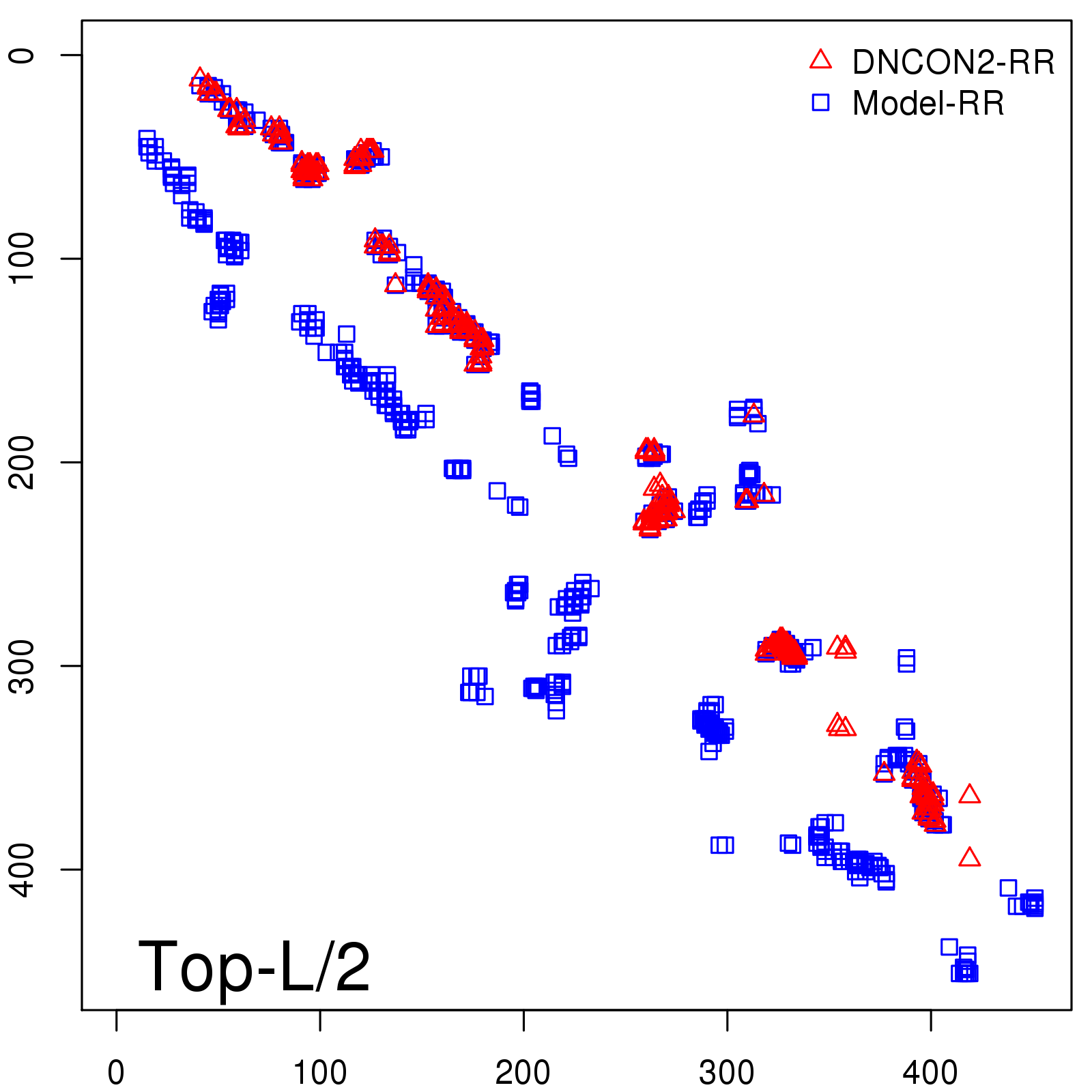
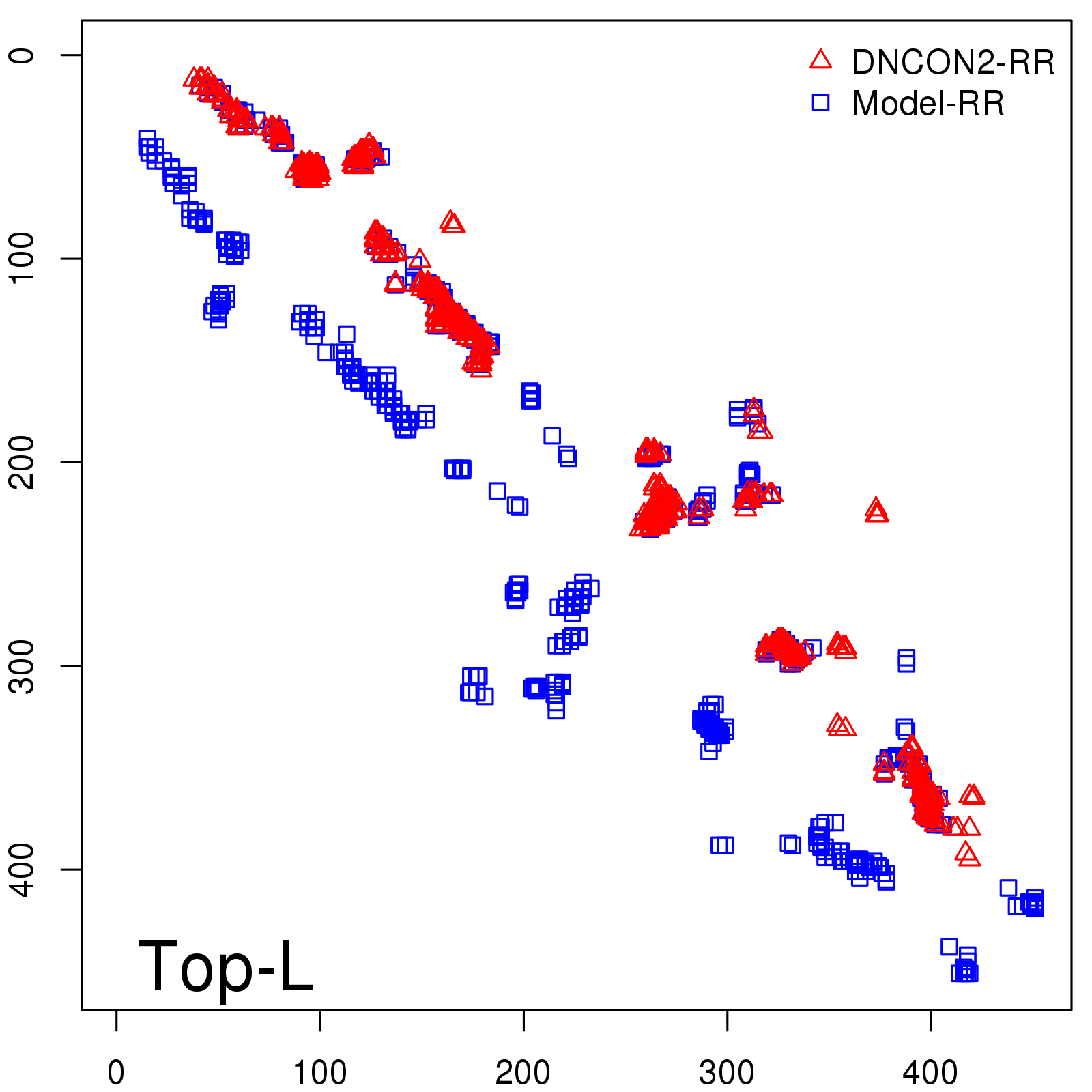
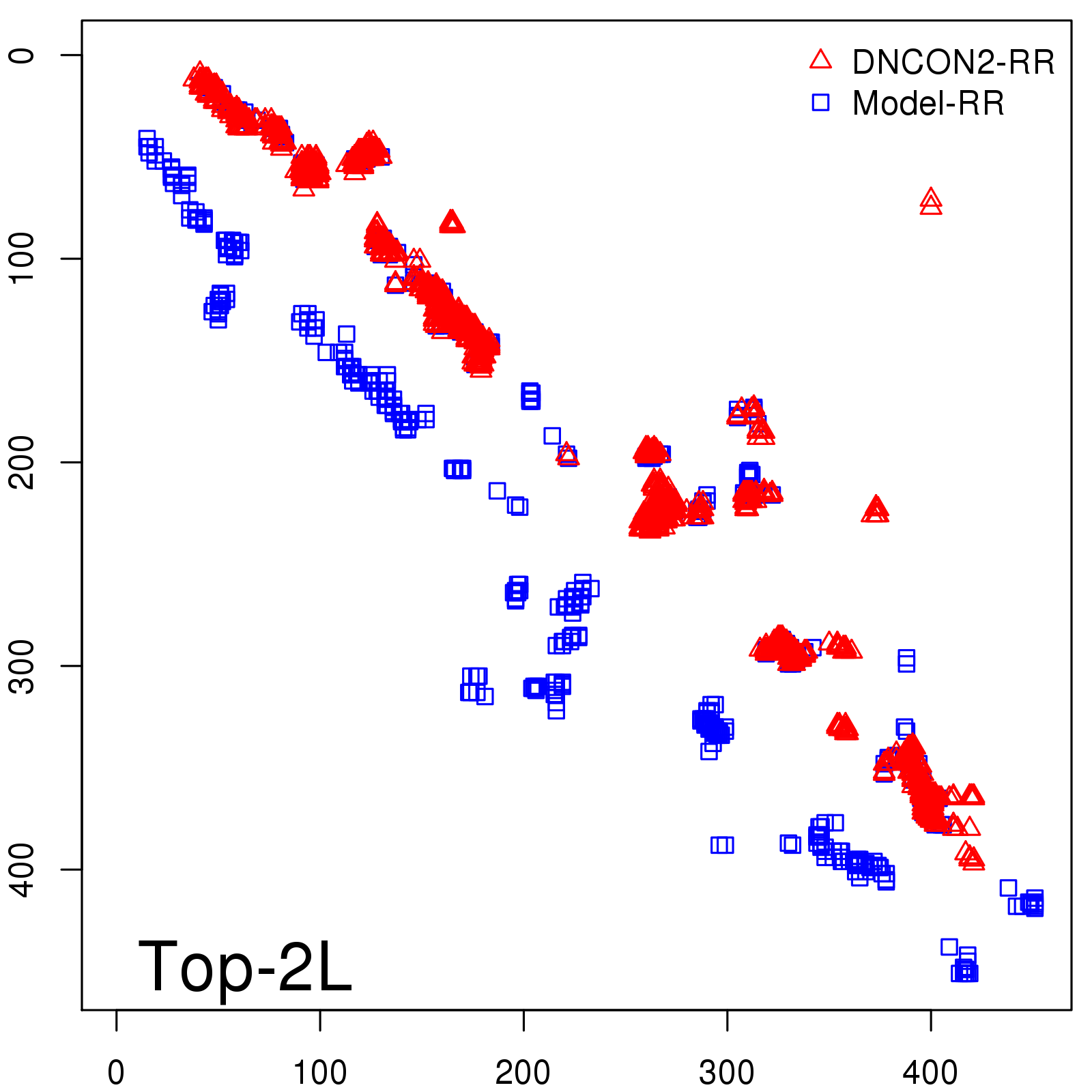
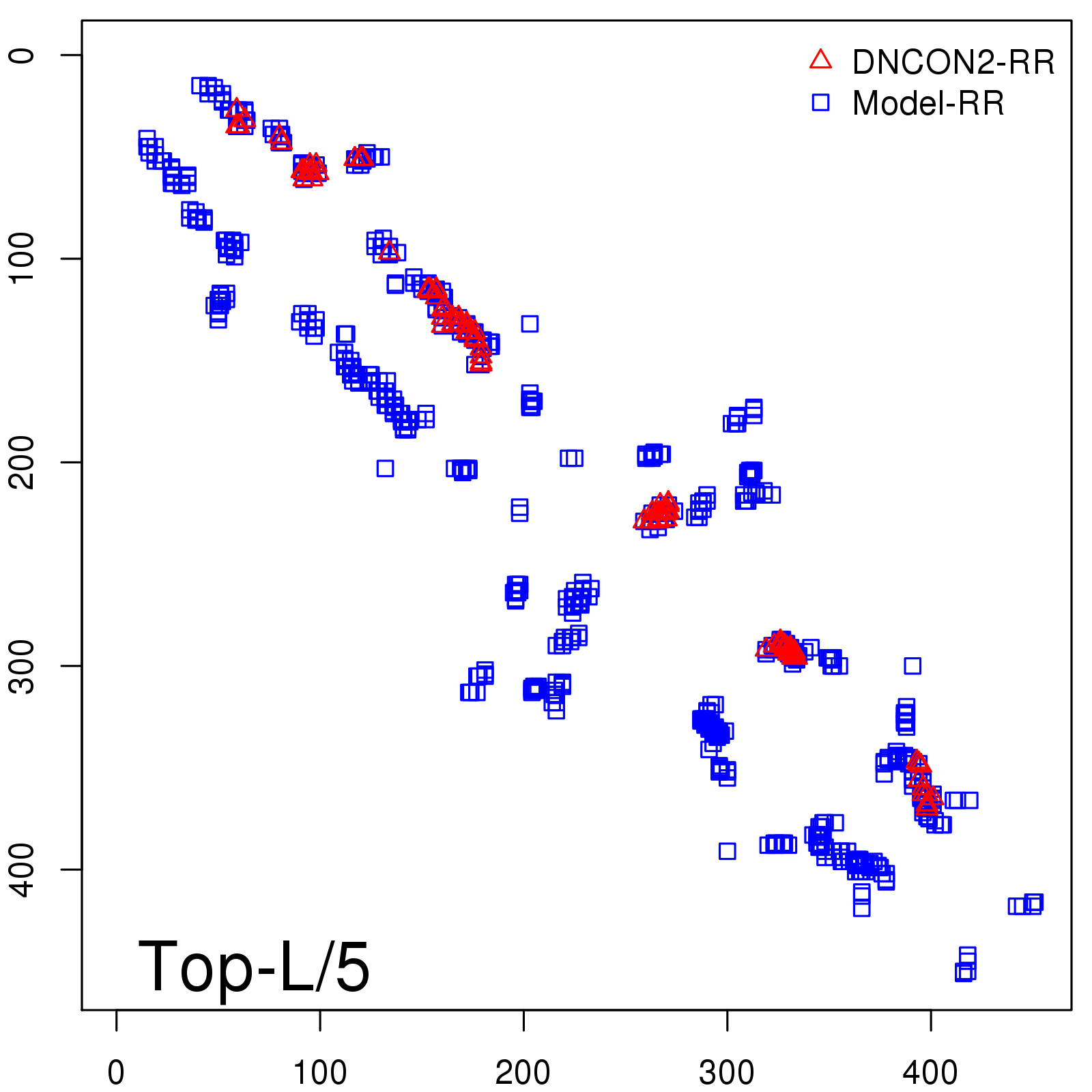
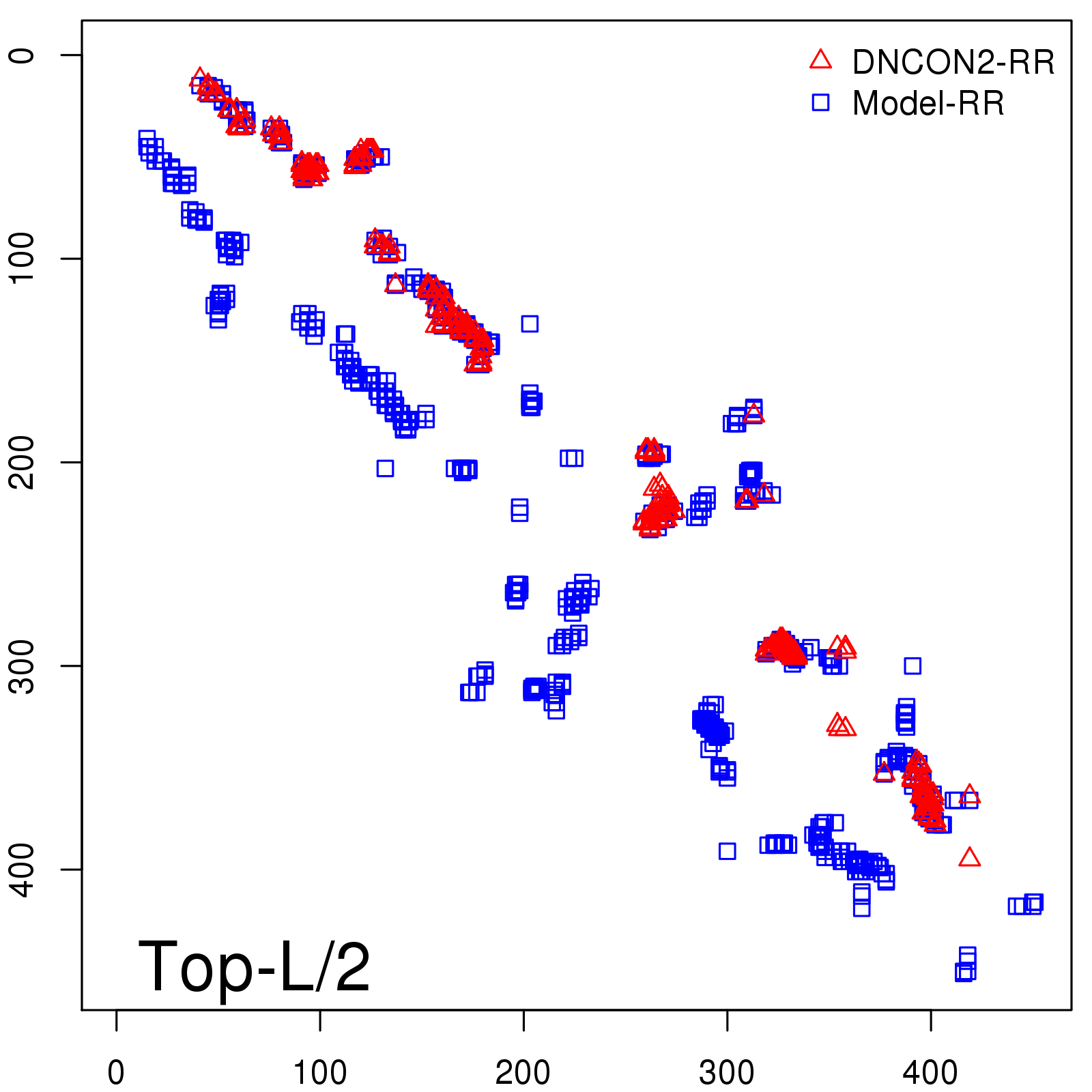
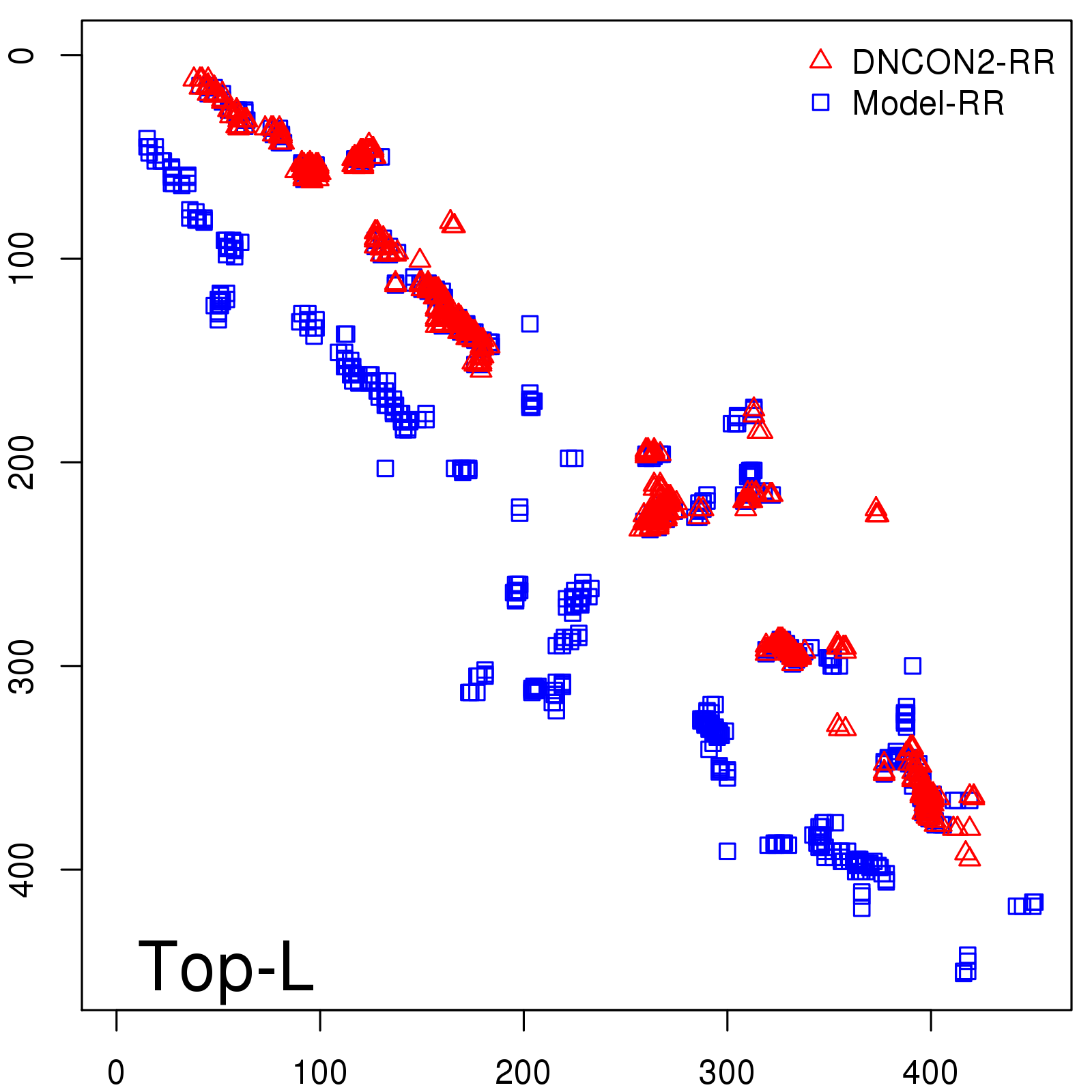
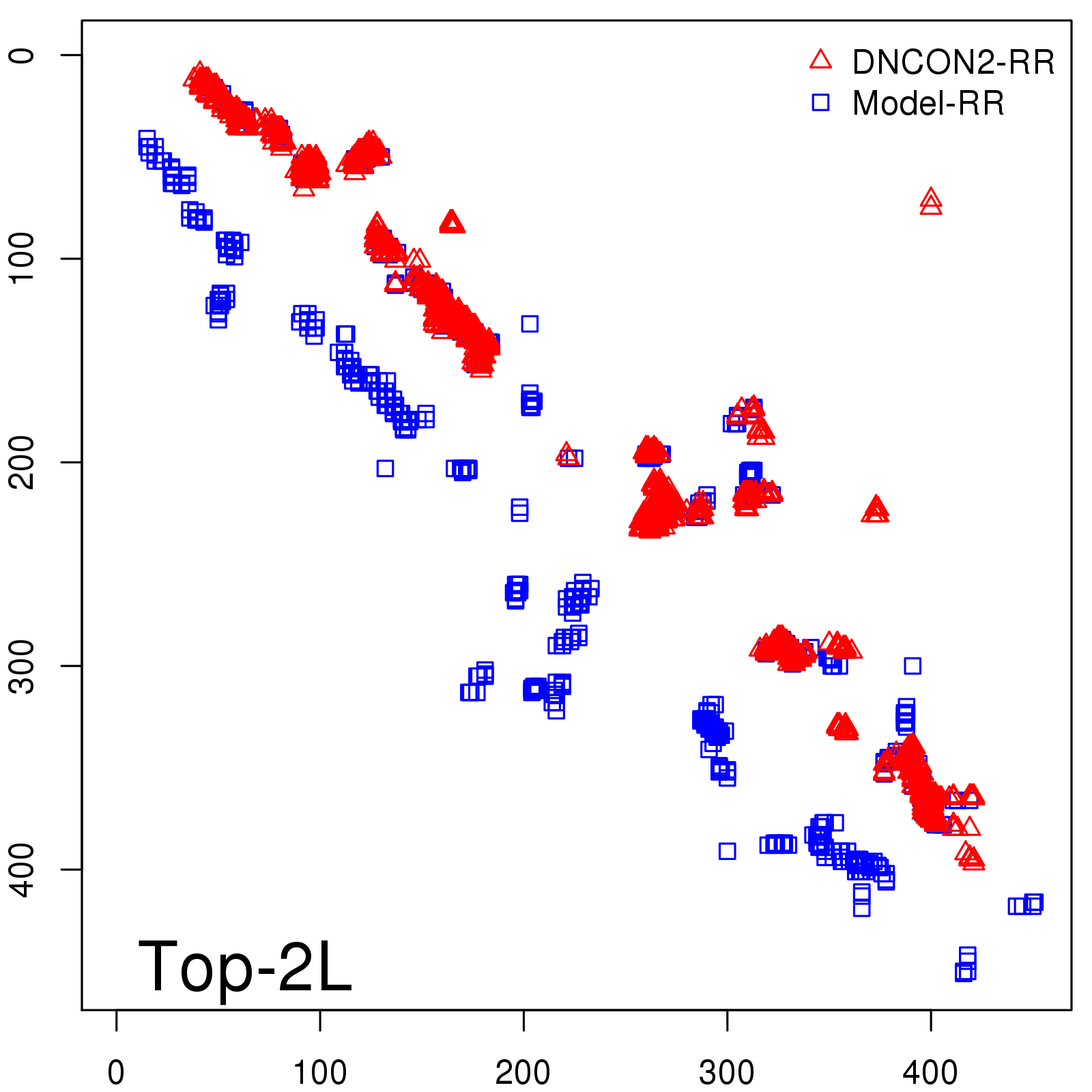
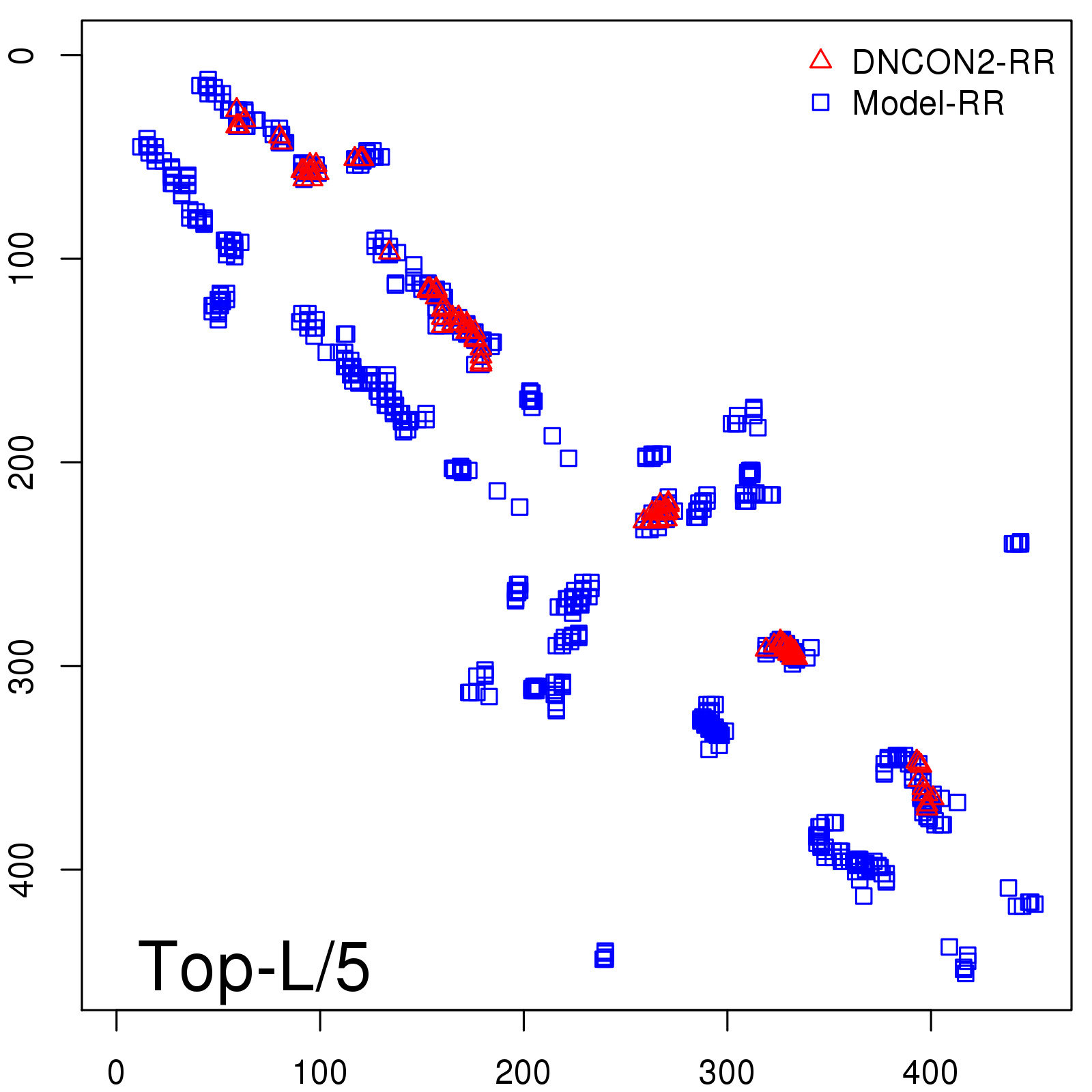
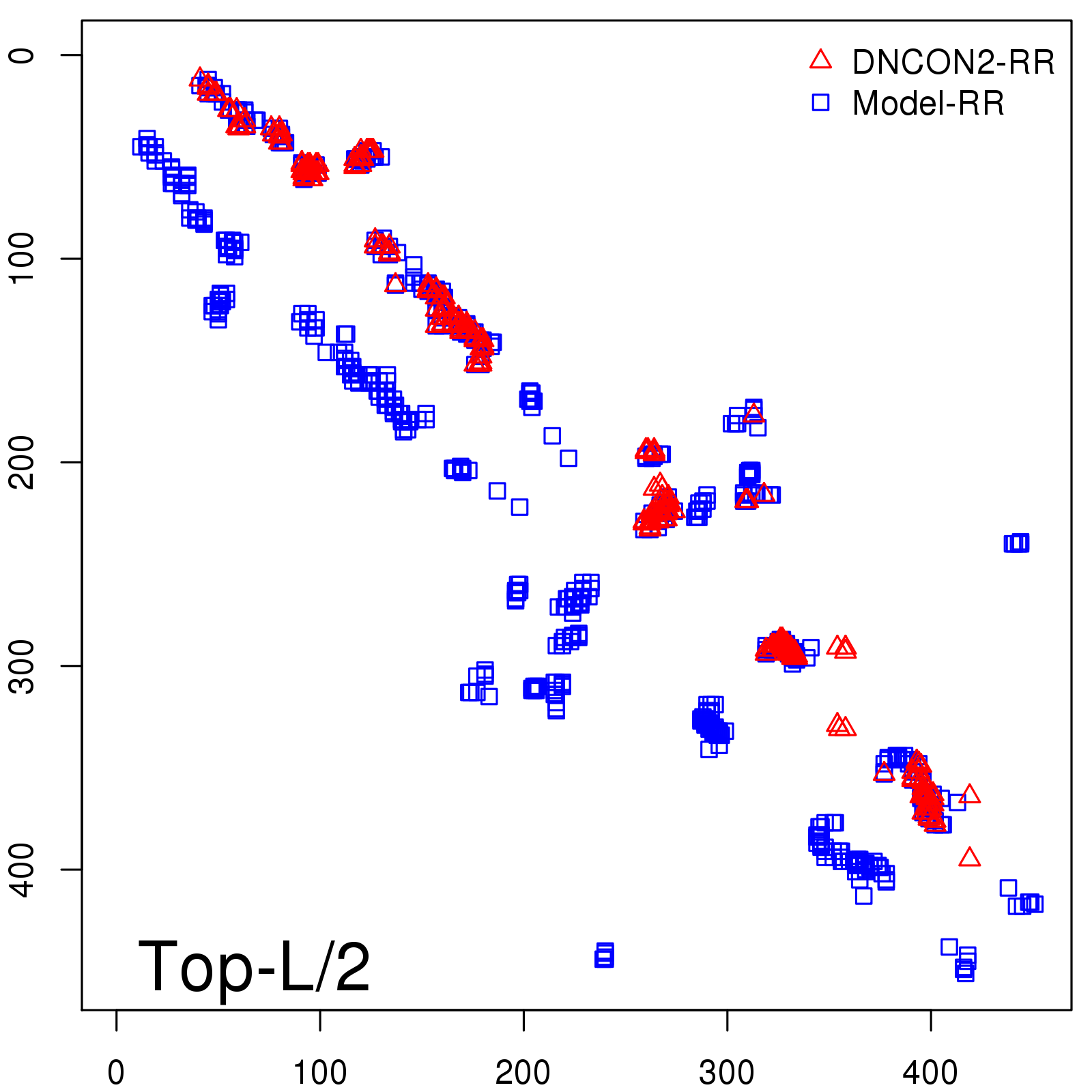
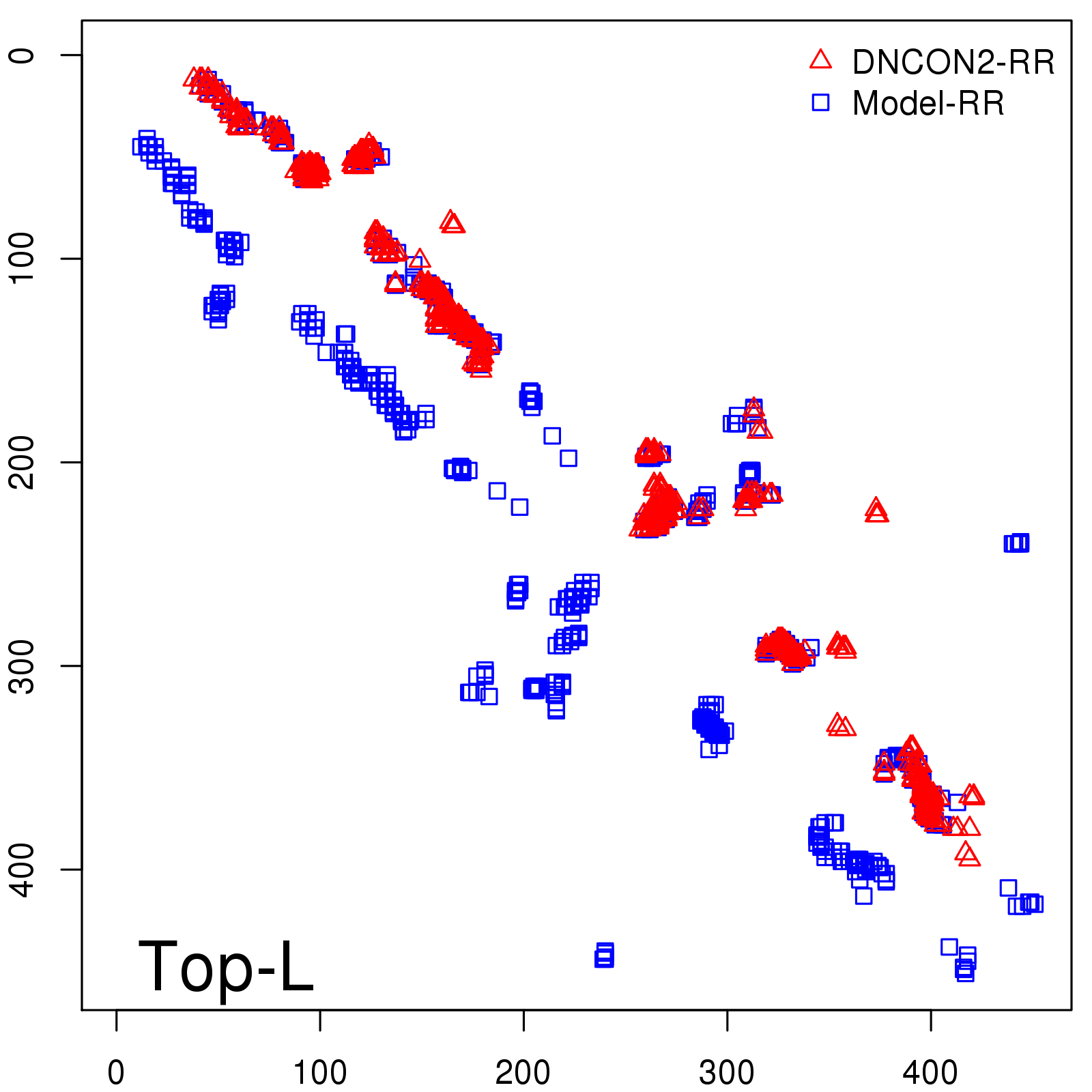
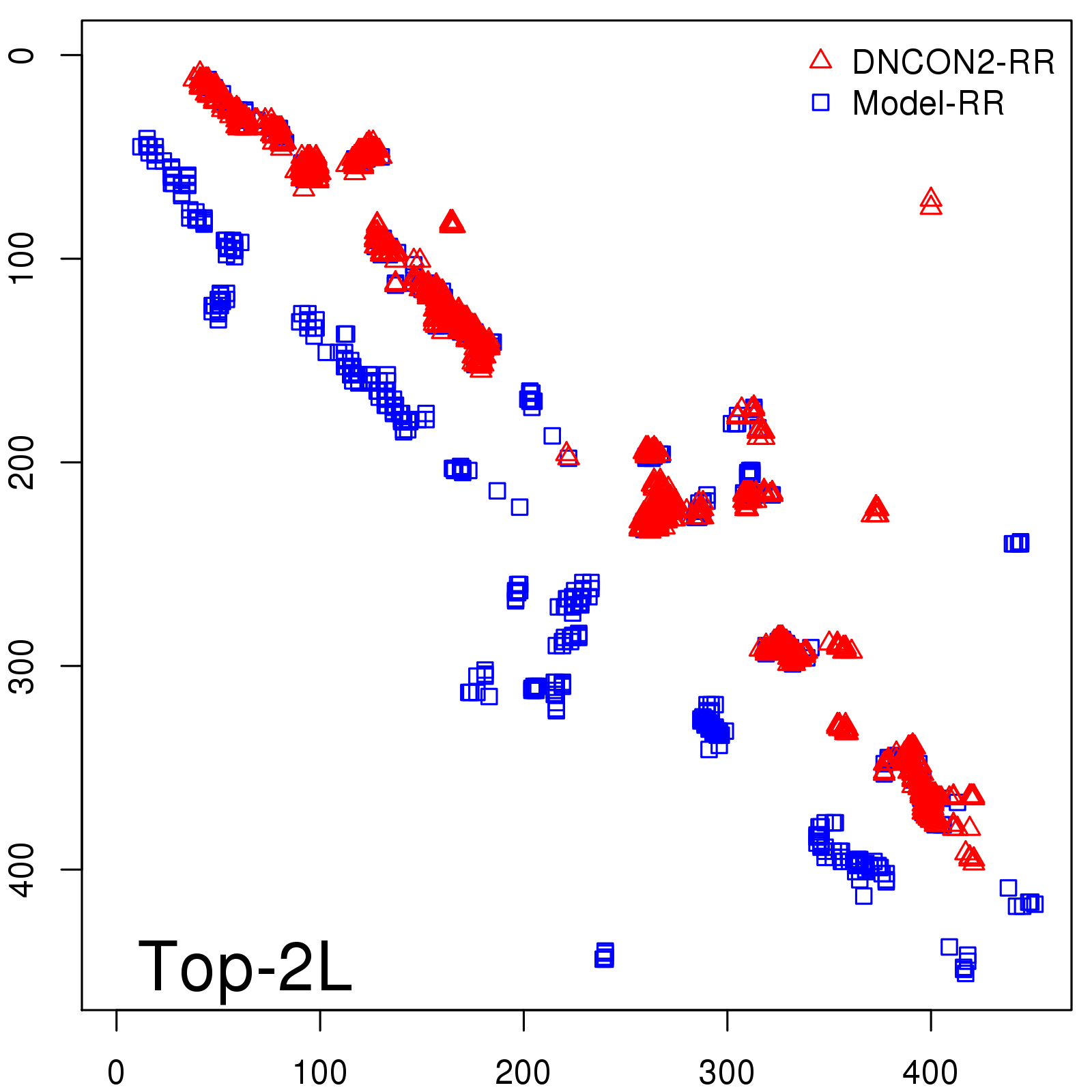
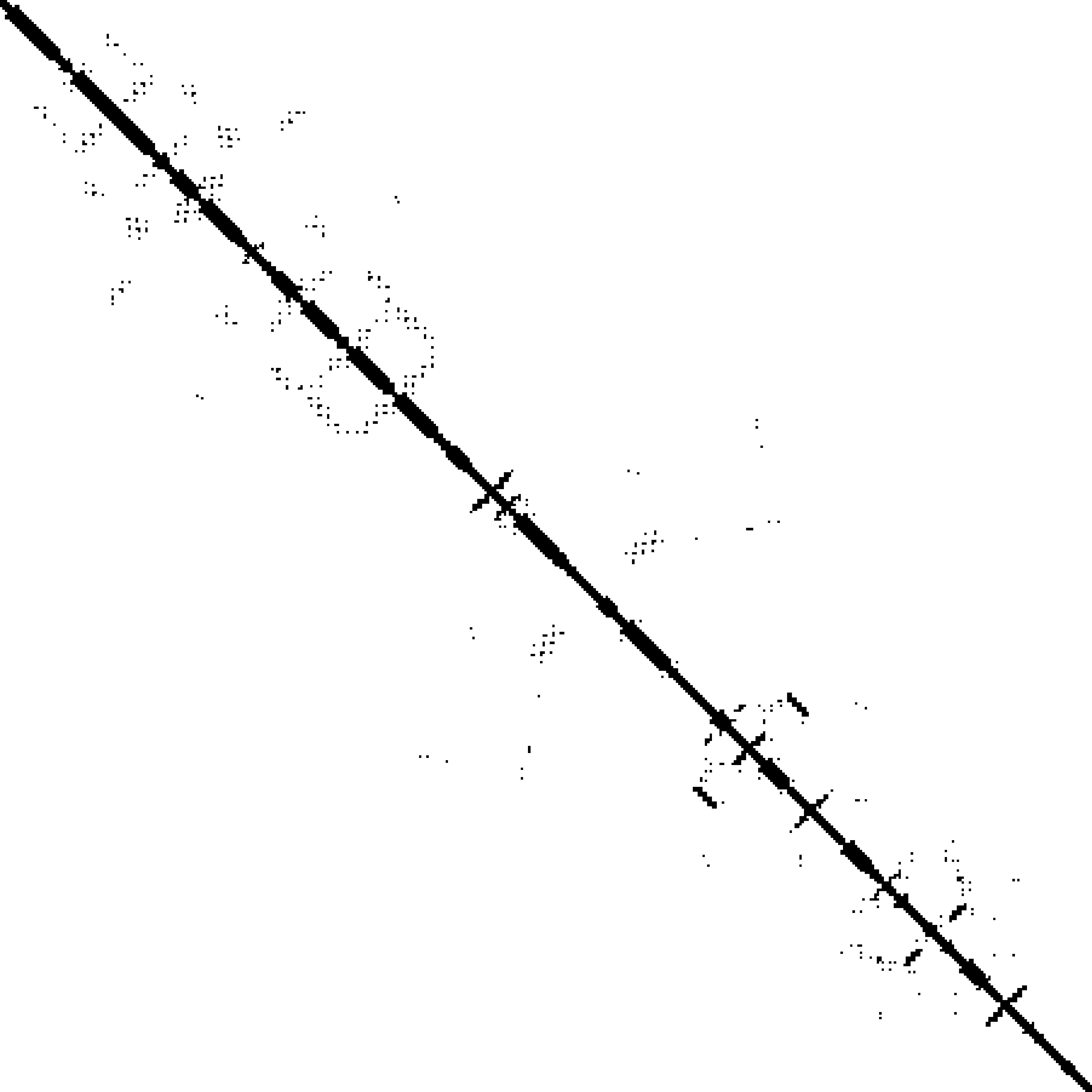
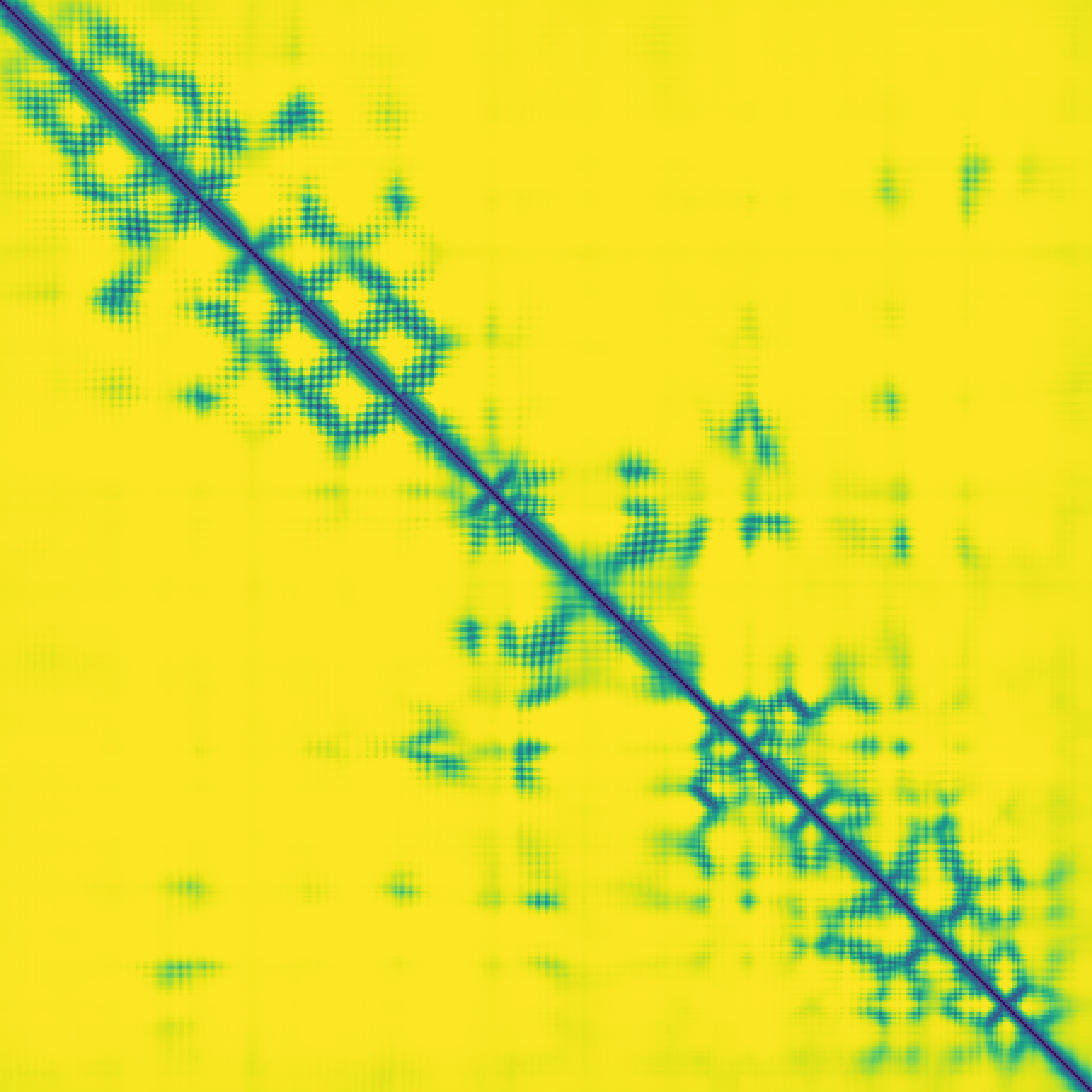
Predicted contact map and distance map
|
Predicted distance map 
|
|
Predicted contact map 
|
Probability to Precision
| Long-Range |
Average probability |
Predicted precision |
| TopL/5 |
0.83 |
82.48 |
| TopL/2 |
0.67 |
70.48 |
Note: The linear model (Average probability vs Predicted precision) was trained on CASP14 dataset
|
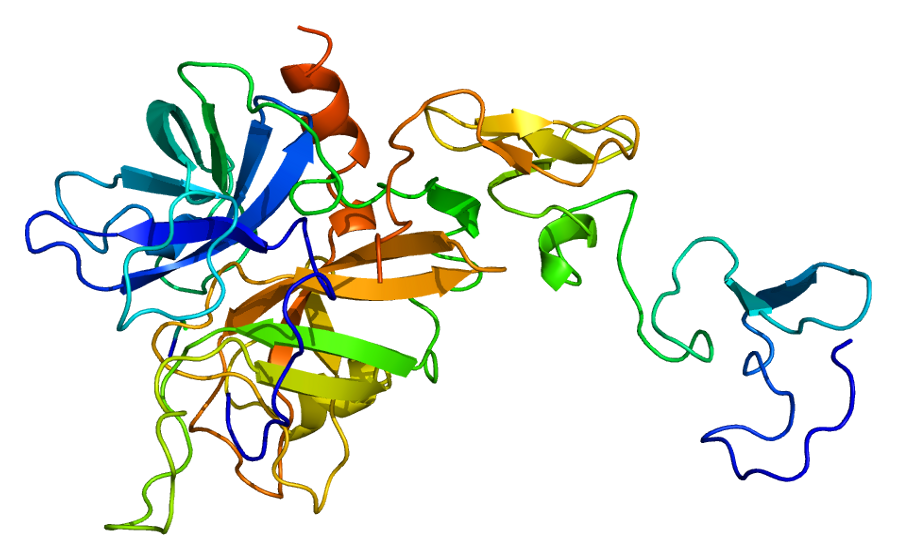
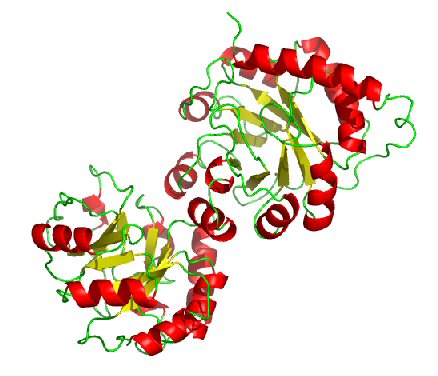
Predicted Top 1 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
94.44 |
| TopL/2 |
76.55 |
| TopL |
50.78 |
| Top2L |
28.60 |
| Alignment |
Number |
| N |
13666 |
| Neff |
2740 |
|
Model comparsion
| Model |
TM-score |
| Model 2 |
0.7216 |
| Model 3 |
0.7369 |
| Model 4 |
0.7324 |
| Model 5 |
0.7207 |
| Average |
0.72790 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 6ez8 |
0.43283 |
| 3j8h |
0.42277 |
| 5kzw |
0.42158 |
| 5kzx |
0.42158 |
| 5yz0 |
0.41827 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 2 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
95.56 |
| TopL/2 |
78.76 |
| TopL |
51.66 |
| Top2L |
28.94 |
| Alignment |
Number |
| N |
13666 |
| Neff |
2740 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.7216 |
| Model 3 |
0.8254 |
| Model 4 |
0.7858 |
| Model 5 |
0.7457 |
| Average |
0.76963 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 4uwa |
0.43836 |
| 4uwe |
0.43533 |
| 5ve8 |
0.42794 |
| 6mfz |
0.42561 |
| 3s4w |
0.42313 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 3 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
96.67 |
| TopL/2 |
80.09 |
| TopL |
50.78 |
| Top2L |
28.60 |
| Alignment |
Number |
| N |
13666 |
| Neff |
2740 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.7369 |
| Model 2 |
0.8254 |
| Model 4 |
0.8390 |
| Model 5 |
0.7509 |
| Average |
0.78805 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5yu6 |
0.44026 |
| 3a6p |
0.43800 |
| 5vlj |
0.43782 |
| 4uwe |
0.43043 |
| 3m63 |
0.42925 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 4 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
94.44 |
| TopL/2 |
76.55 |
| TopL |
51.22 |
| Top2L |
28.60 |
| Alignment |
Number |
| N |
13666 |
| Neff |
2740 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.7324 |
| Model 2 |
0.7858 |
| Model 3 |
0.8390 |
| Model 5 |
0.7132 |
| Average |
0.76760 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5luq |
0.44243 |
| 5gl1 |
0.43353 |
| 5gl0 |
0.43237 |
| 6mu2 |
0.43230 |
| 6jiu |
0.42922 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
Predicted Top 5 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
95.56 |
| TopL/2 |
79.65 |
| TopL |
53.22 |
| Top2L |
30.04 |
| Alignment |
Number |
| N |
13666 |
| Neff |
2740 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.7207 |
| Model 2 |
0.7457 |
| Model 3 |
0.7509 |
| Model 4 |
0.7132 |
| Average |
0.73263 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 4zhj |
0.44533 |
| 4rh7 |
0.43995 |
| 6or5 |
0.43825 |
| 6jxa |
0.43474 |
| 6rla |
0.43299 |
|
Note: This is multi-domain structure, check alignments and domain qualities in individual domains!
|
References
[1] Hou, J., Wu, T., Cao, R., & Cheng, J. (2019). Protein tertiary structure modeling driven by deep learning and contact distance prediction in CASP13. bioRxiv, 552422.
[2] Li, J., Deng, X., Eickholt, J., & Cheng, J. (2013). Designing and benchmarking the MULTICOM protein structure prediction system. BMC structural biology, 13(1), 2.
[3] Cheng, J., Li, J., Wang, Z., Eickholt, J., & Deng, X. (2012). The MULTICOM toolbox for protein structure prediction. BMC bioinformatics, 13(1), 65.
[4] Wang, Z., Eickholt, J., & Cheng, J. (2010). MULTICOM: a multi-level combination approach to protein structure prediction and its assessments in CASP8. Bioinformatics, 26(7), 882-888.
 )
) )
)

 )
) )
) )
) )
) )
)