Q6F8C5
multicom
Q6F8C5
full_length
Q6F8C5
Results of Structure Prediction for Target Name: Q6F8C5 ( Click  )
)
Domain Boundary prediction ( View  )
)

>Q6F8C5: 1-172
| 1-60: |
M | L | A | F | N | L | L | R | S | A | H | M | L | N | P | A | L | Q | V | L | A | K | D | G | S | I | A | D | N | T | Y | Q | P | I | A | E | D | G | V | I | P | A | G | D | V | V | M | T | T | A | Q | L | D | Q | L | T | A | I | Q | G |
| 61-119: |
K | K | A | L | Y | I | T | V | N | D | S | P | E | T | H | Q | F | P | L | D | Q | L | D | A | I | F | I | E | F | A | G | F | N | D | G | R | G | Y | S | F | A | A | L | L | R | R | Q | G | Y | Q | G | E | L | R | A | V | G | D | I | F |
| 121-172: |
K | D | V | L | N | Y | L | K | R | S | G | F | D | T | F | V | I | K | E | G | K | D | I | Q | E | A | A | A | G | L | N | D | F | R | Q | P | Y | Q | A | S | T | A | V | P | Q | A | D | Y | Q | T | G | A |
| 1-60: |
C | C | C | H | H | H | H | C | C | C | C | C | C | C | C | C | C | H | H | H | H | C | C | C | C | E | E | C | C | C | E | E | E | E | C | C | C | C | C | C | C | C | C | C | E | E | E | E | H | H | H | H | H | H | H | H | H | C | C | C |
| 61-119: |
C | E | E | E | E | E | C | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C | C | E | E | E | E | E | C | C | C | C | C | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C | C | C | C | E | E | E | E | C | C | H | H | H |
| 121-172: |
H | H | H | H | H | H | H | H | H | C | C | C | C | E | E | E | E | C | C | C | C | C | H | H | H | H | H | H | H | H | H | C | C | C | C | H | E | C | C | C | C | C | C | C | C | H | H | H | H | H | C | C |
|
| | H(Helix): 62(36.05%) | E(Strand): 29(16.86%) | C(Coil): 81(47.09%) |
| 1-60: |
E | B | B | B | E | B | B | E | B | E | E | B | B | B | E | B | B | E | E | B | B | E | E | E | E | B | B | E | B | E | B | B | B | B | E | E | E | E | E | B | E | E | E | E | B | B | B | B | B | E | B | B | E | E | B | E | E | E | E | E |
| 61-119: |
E | B | B | B | B | B | E | B | E | E | E | B | E | E | B | B | E | B | B | E | E | B | B | B | B | B | B | B | B | E | E | B | E | E | B | B | B | B | B | B | B | B | B | B | B | E | B | E | B | E | B | B | B | B | B | E | B | E | B | B |
| 121-172: |
B | B | B | B | B | B | B | B | B | B | B | B | E | B | B | E | B | E | E | E | E | E | B | E | E | B | B | E | B | B | E | E | B | E | E | E | B | B | E | E | B | E | E | E | E | B | B | B | E | E | E | E |
|
| | e(Exposed): 75(43.6%) | b(Buried): 97(56.4%) |
| 1-60: |
T | T | T | T | T | T | T | T | T | T | T | T | T | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 61-119: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N |
| 121-172: |
N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | N | T | T | T | T |
|
| | N(Normal): 155(90.12%) | T(Disorder): 17(9.88%) |
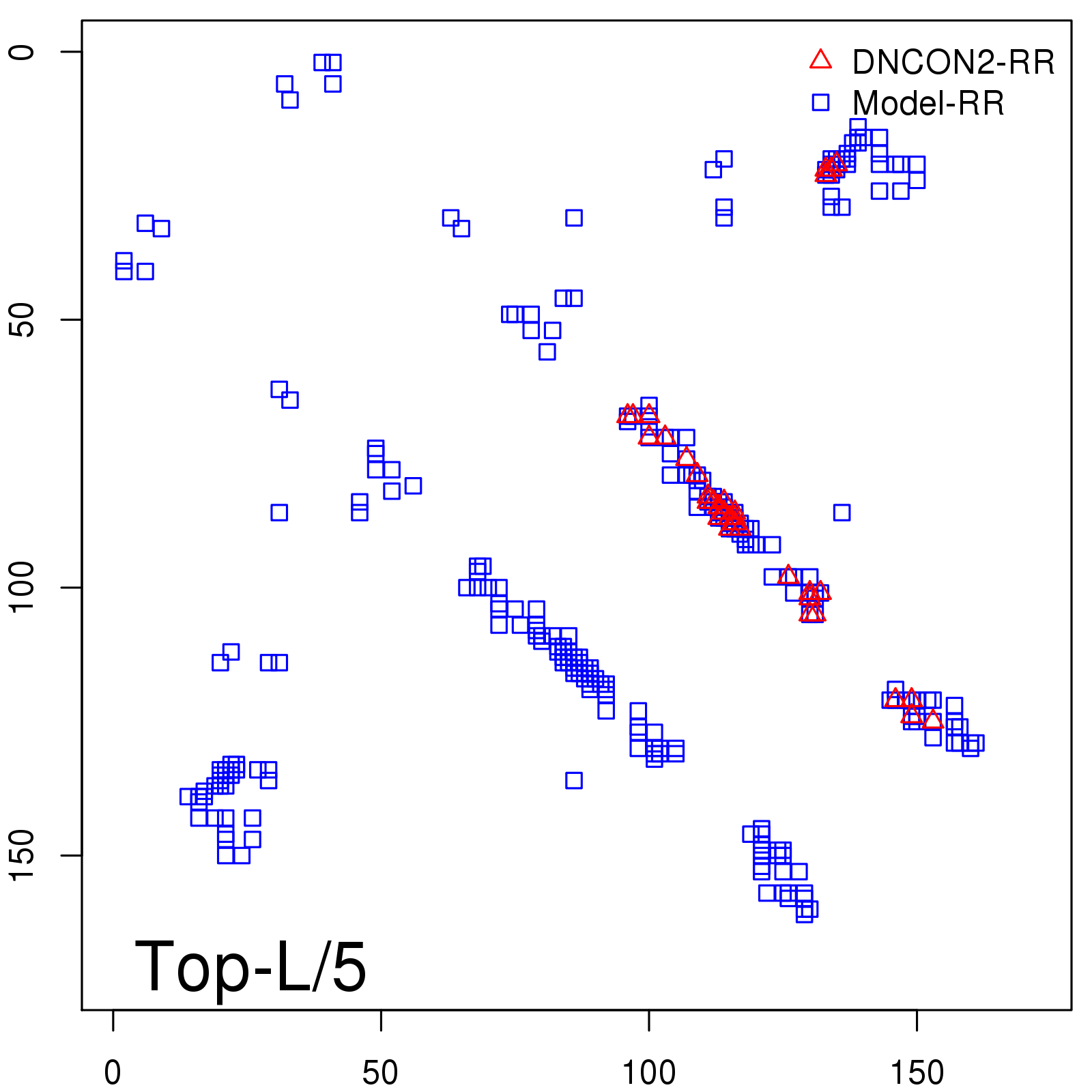
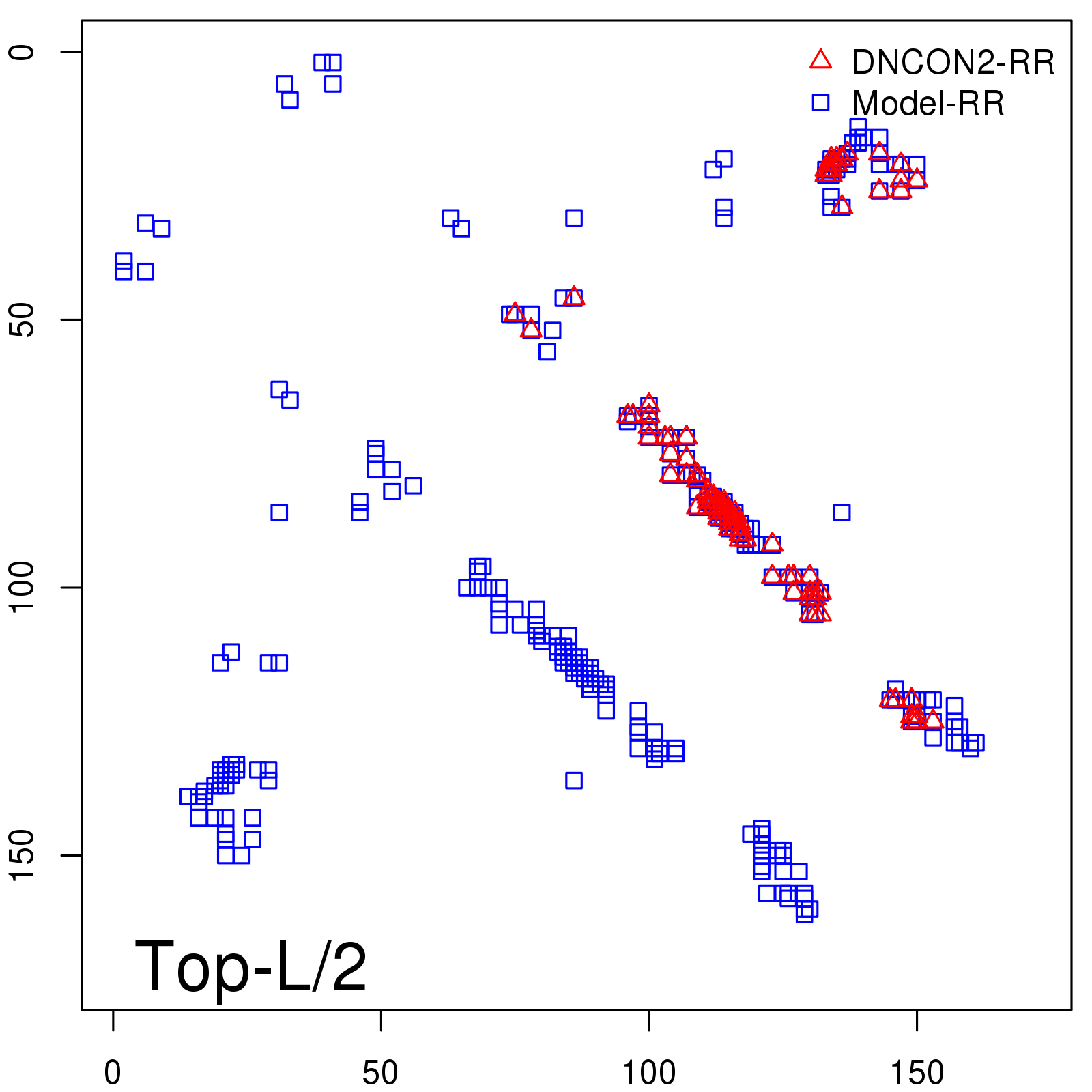
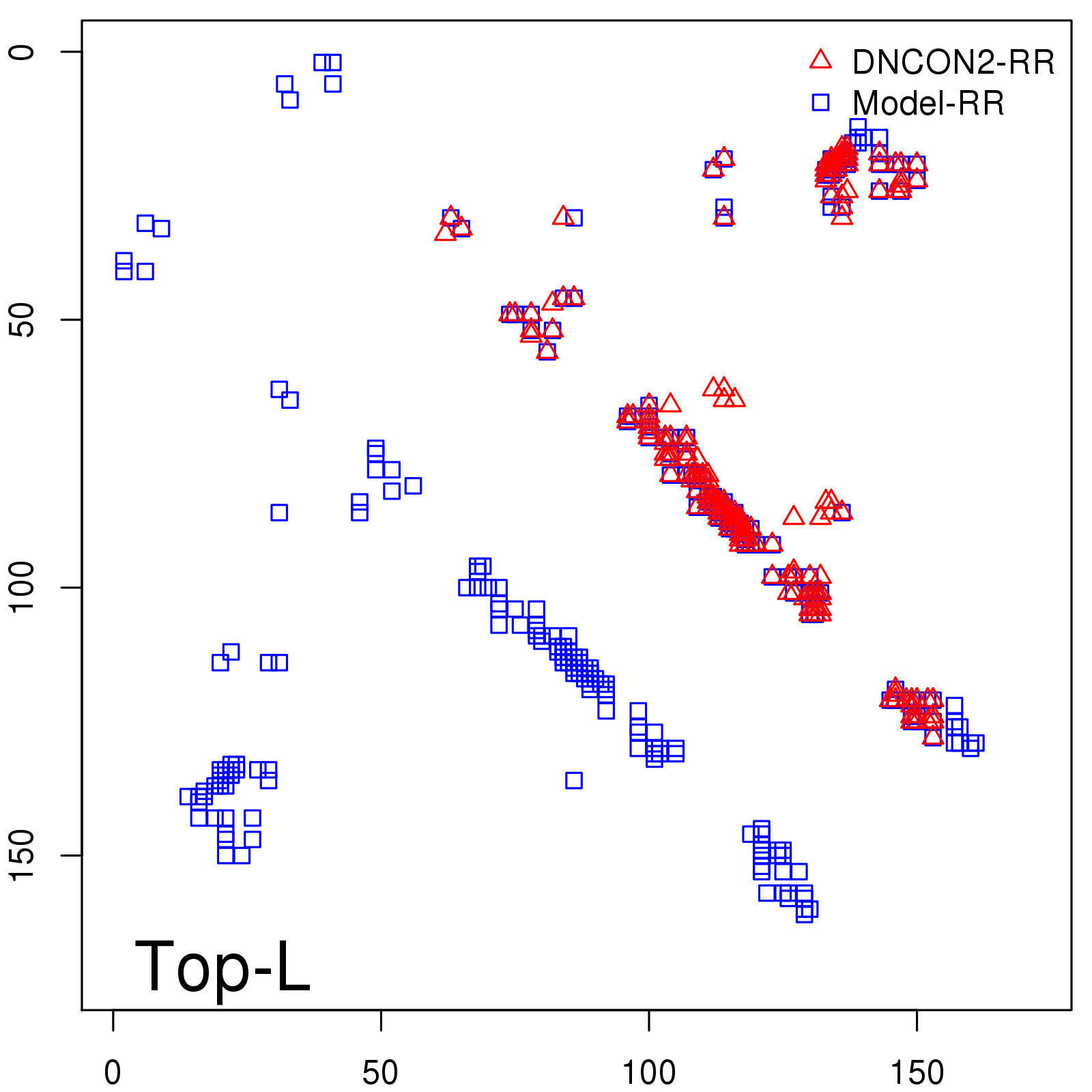
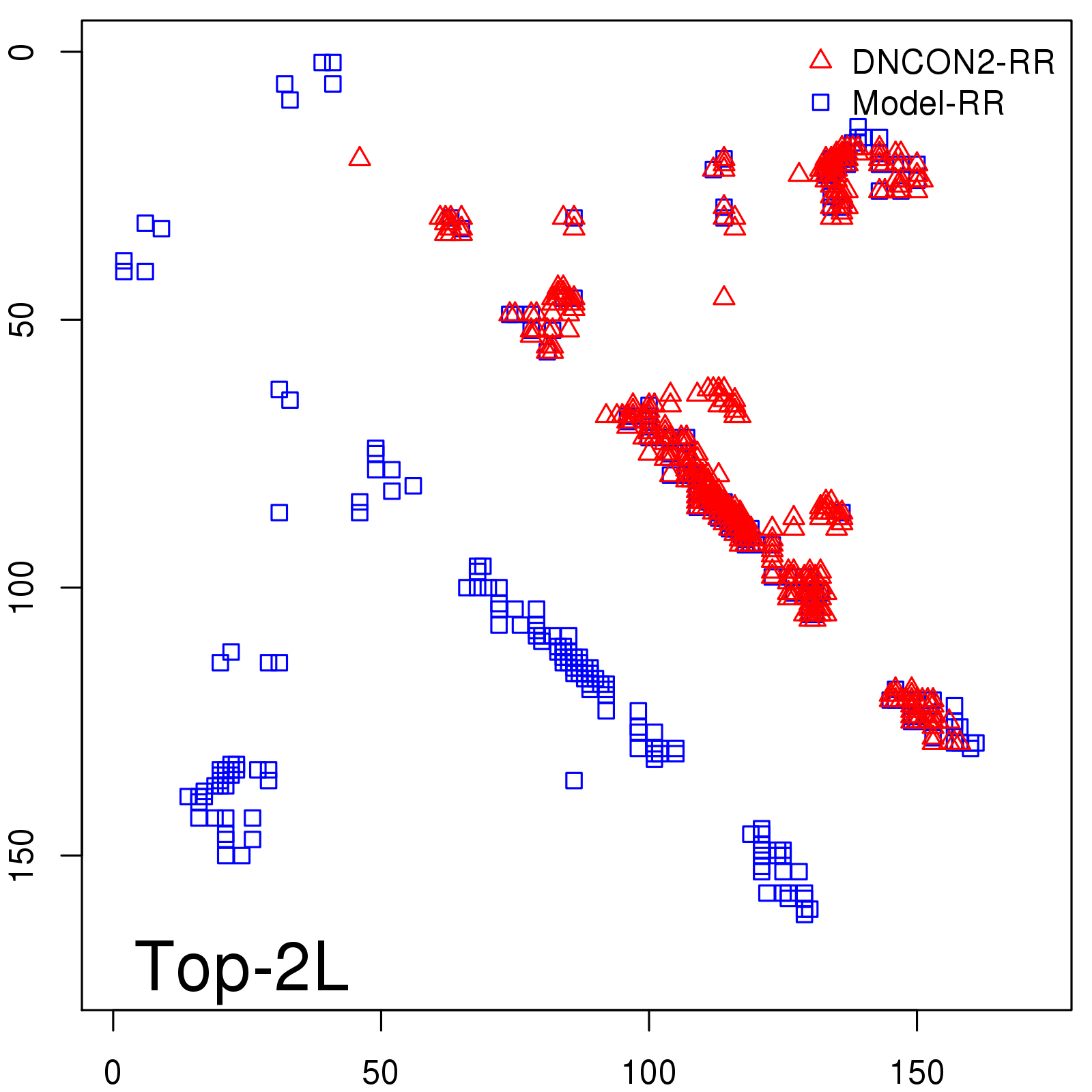
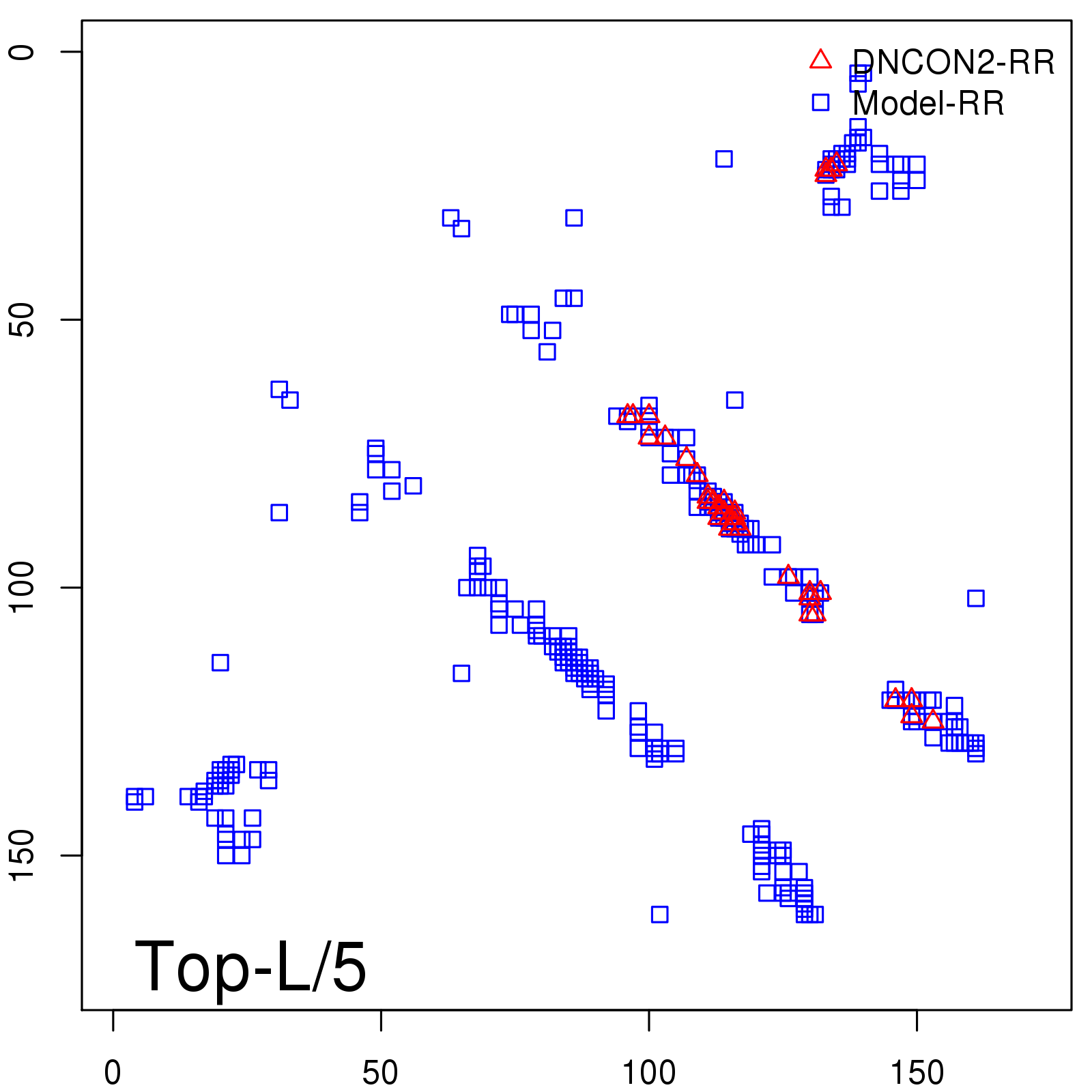
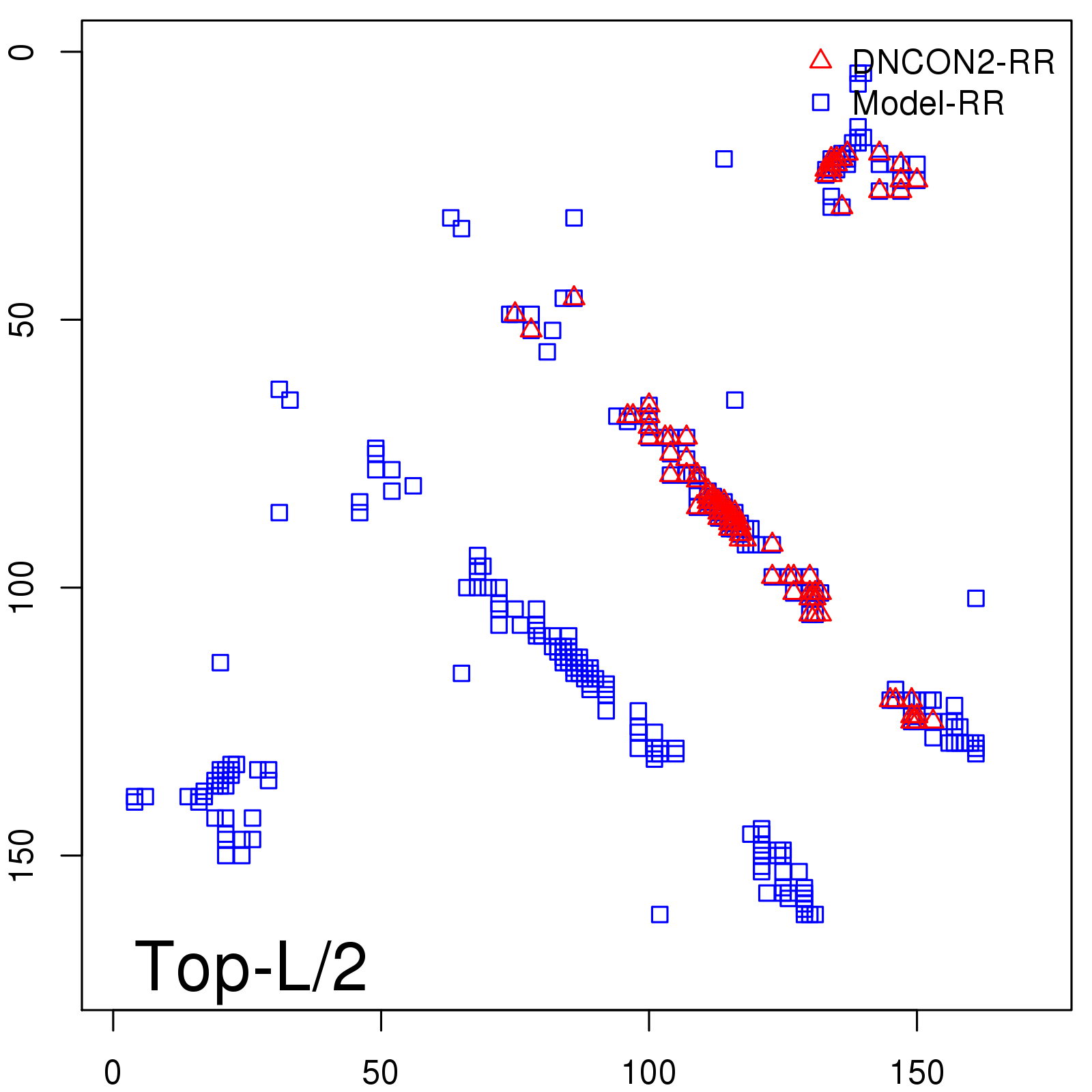
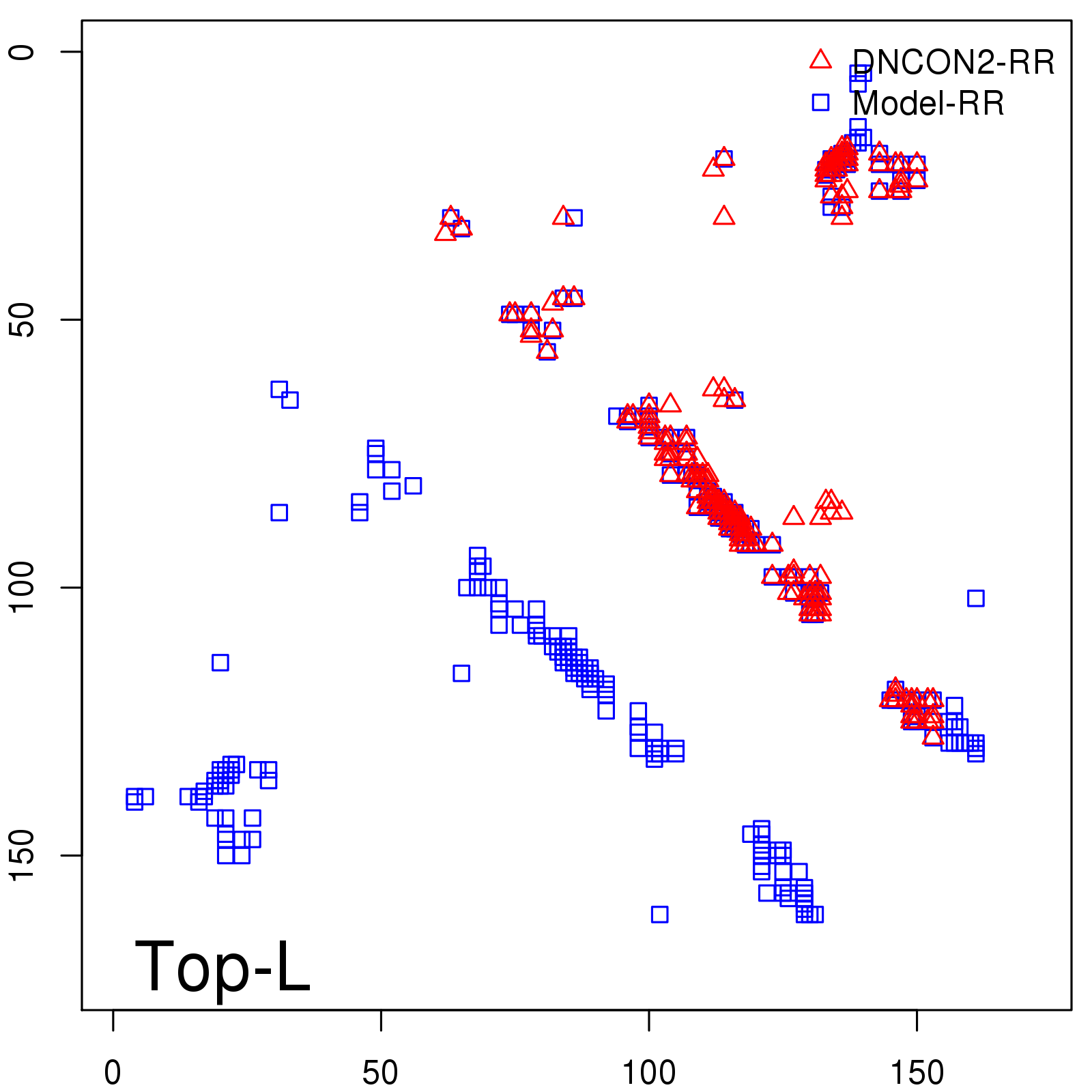
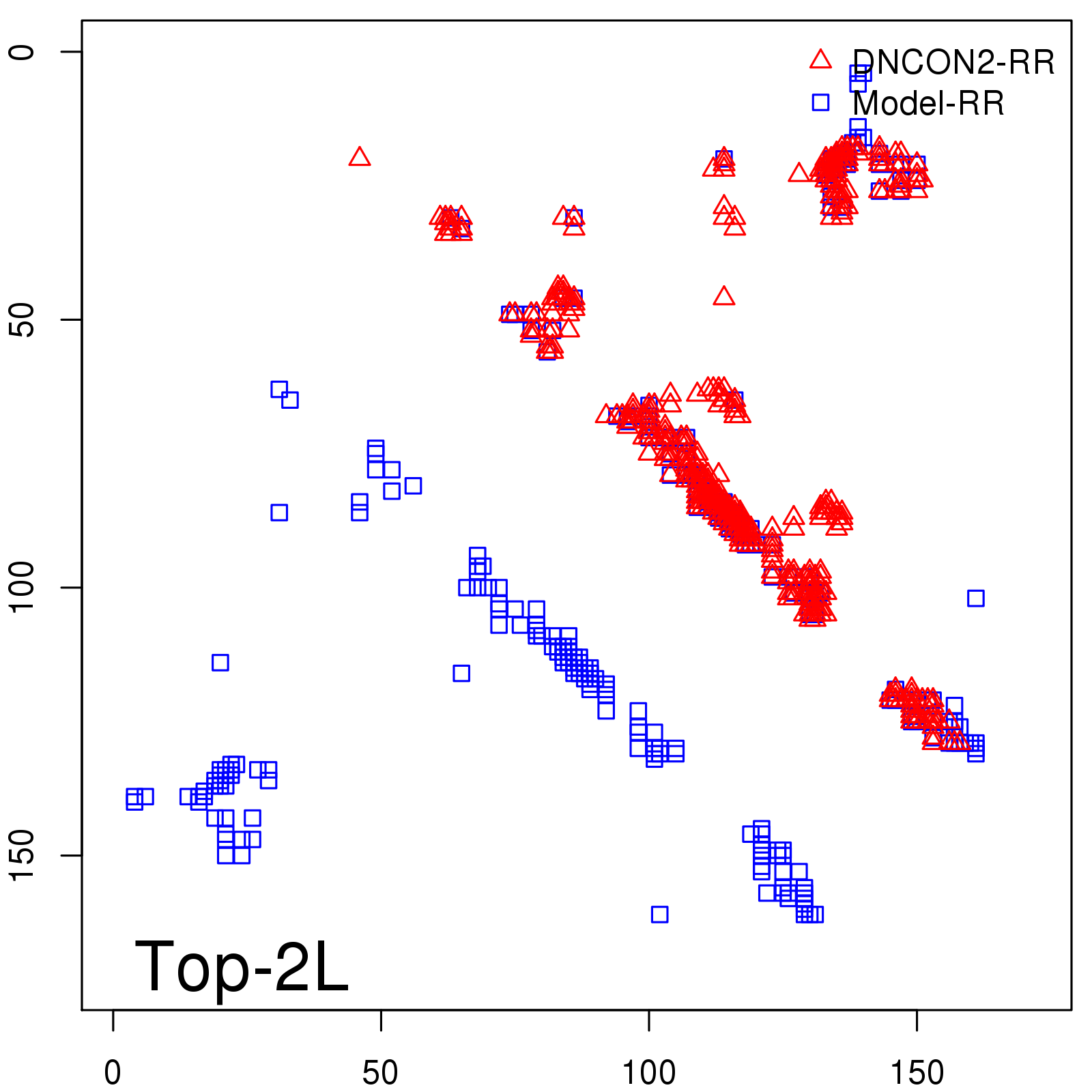
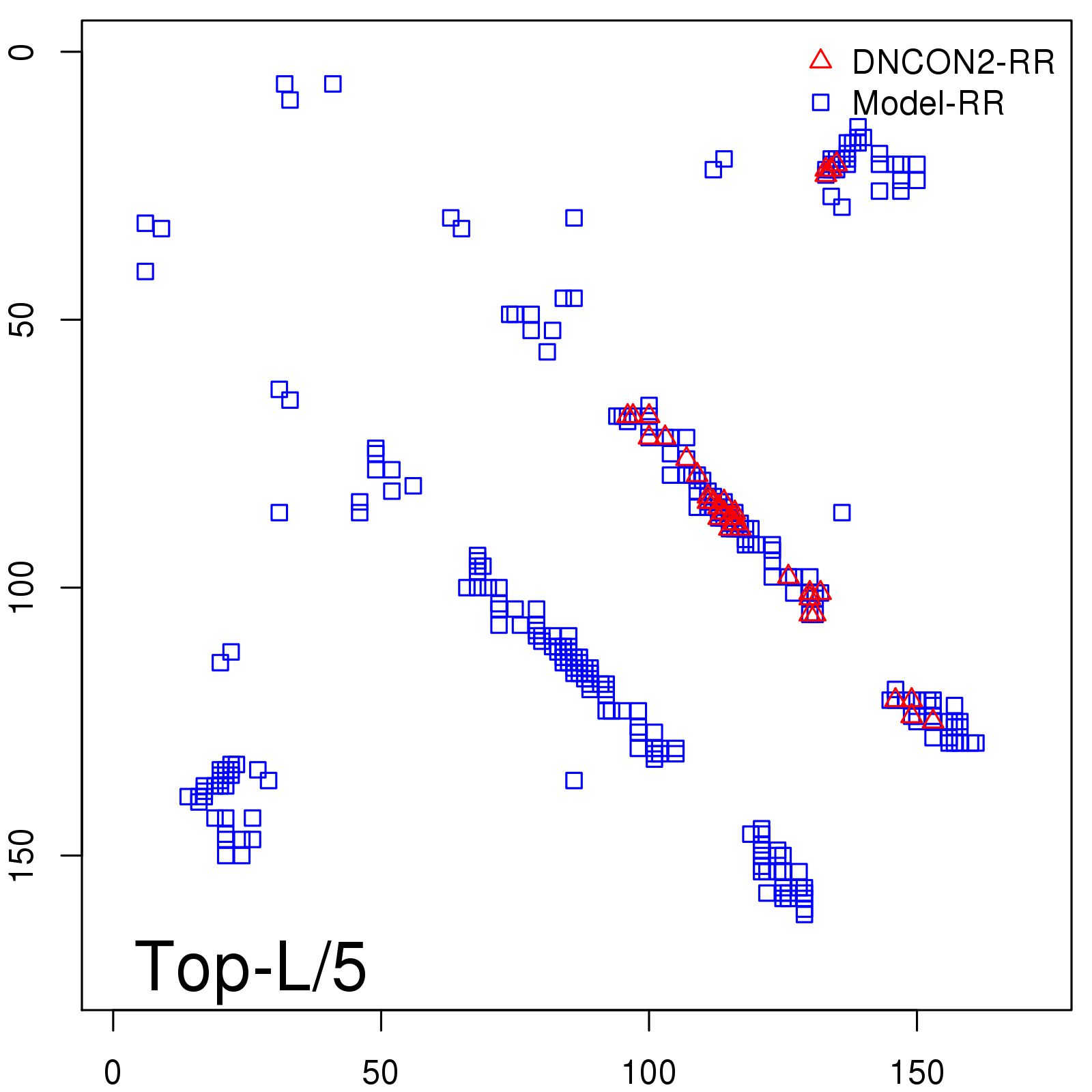
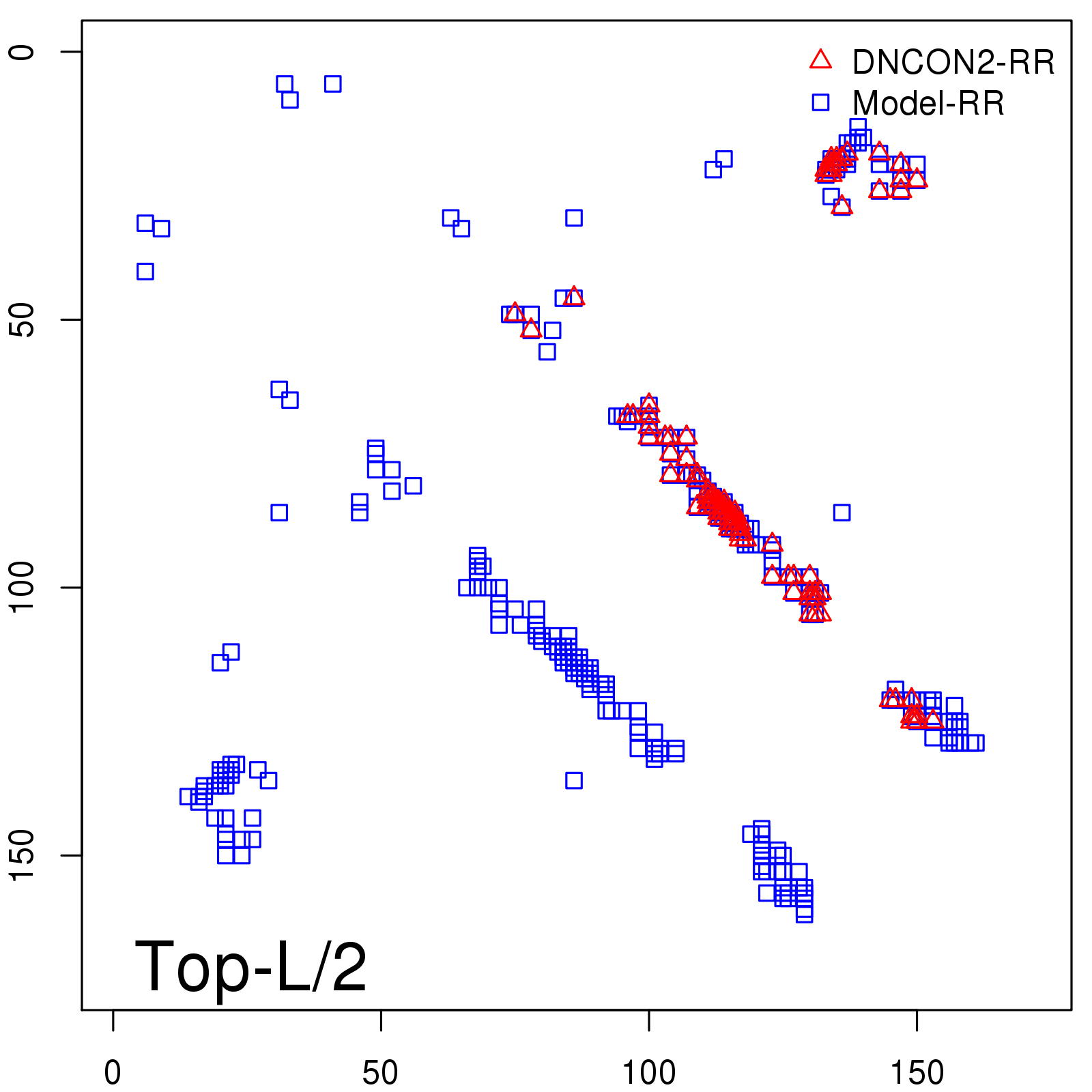
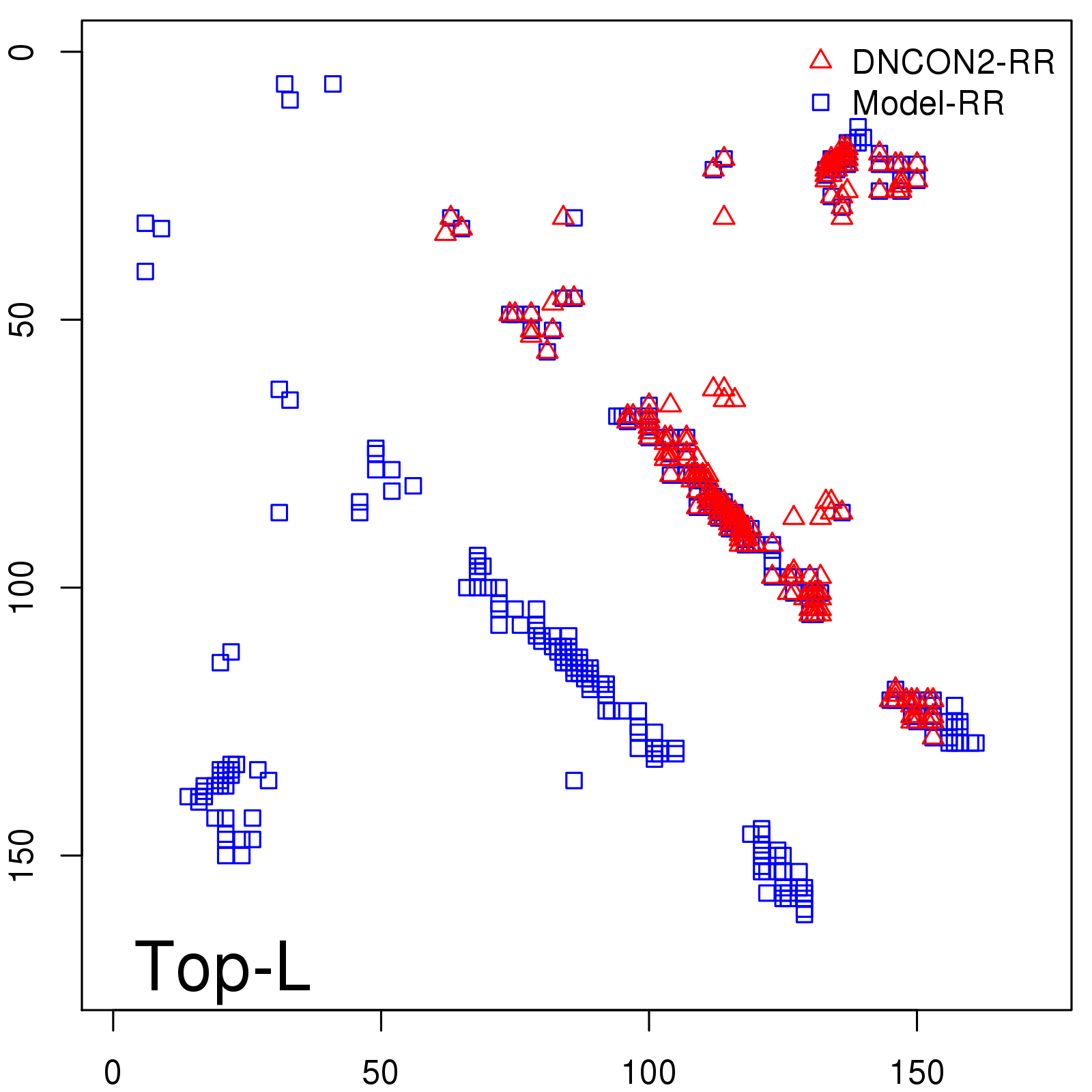
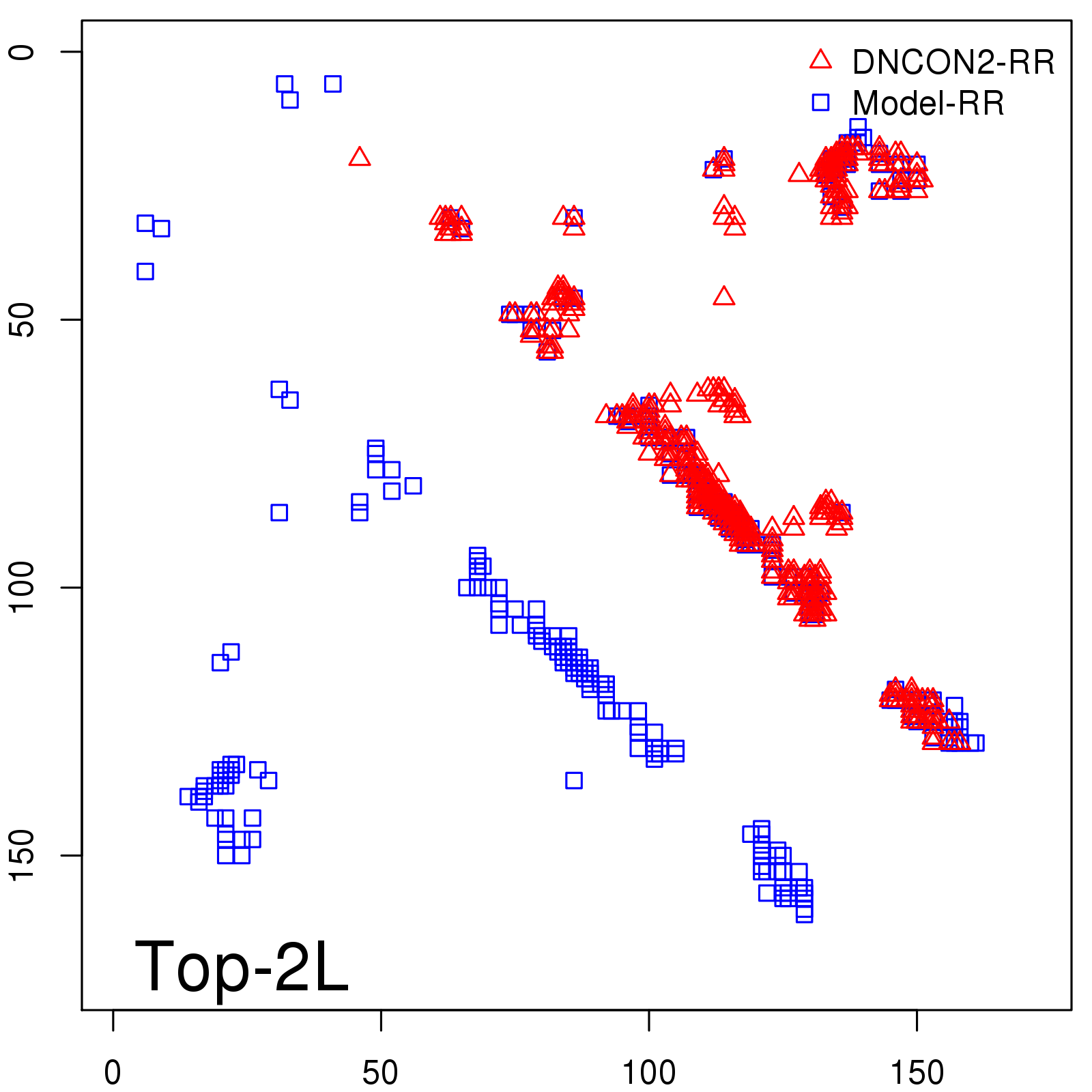
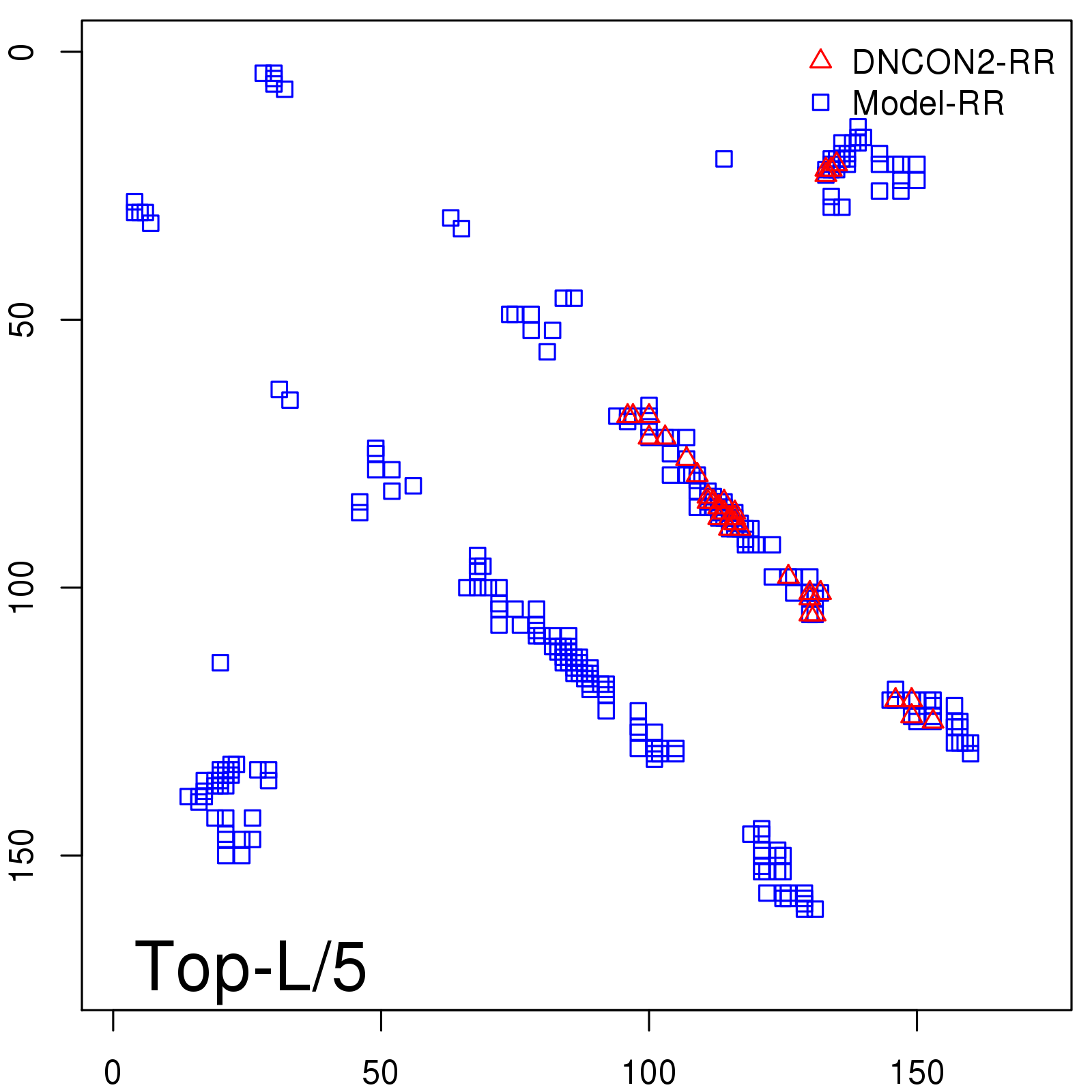
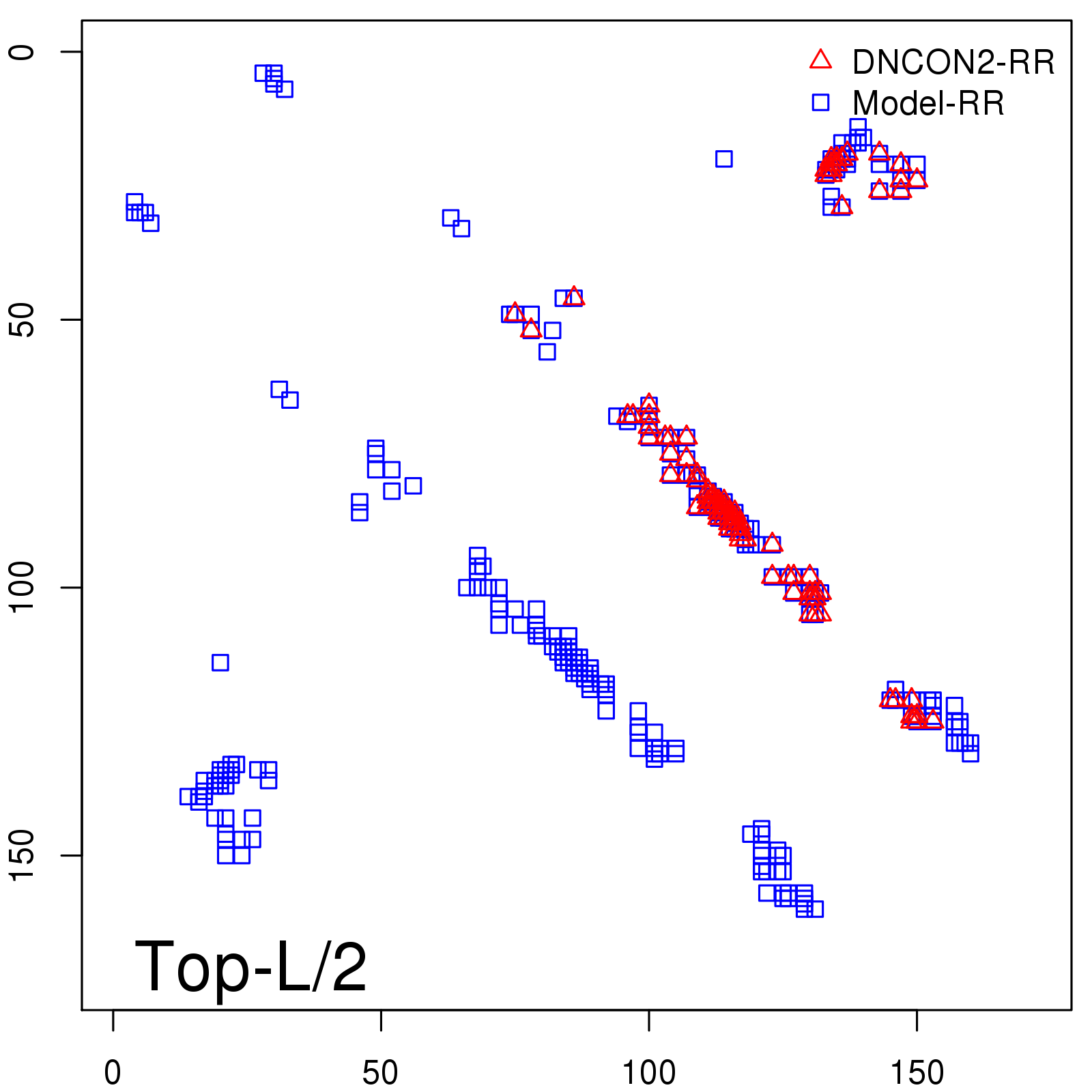
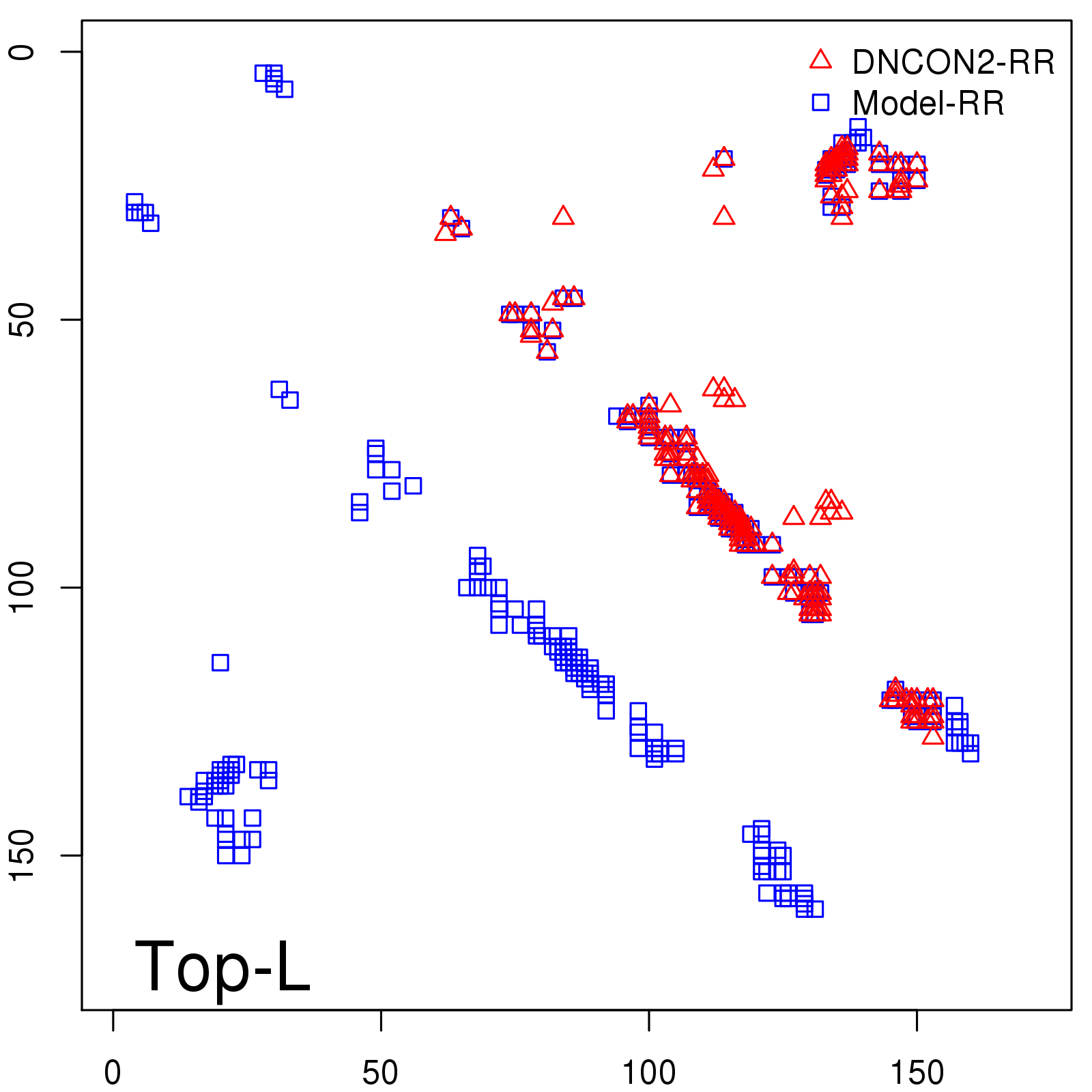
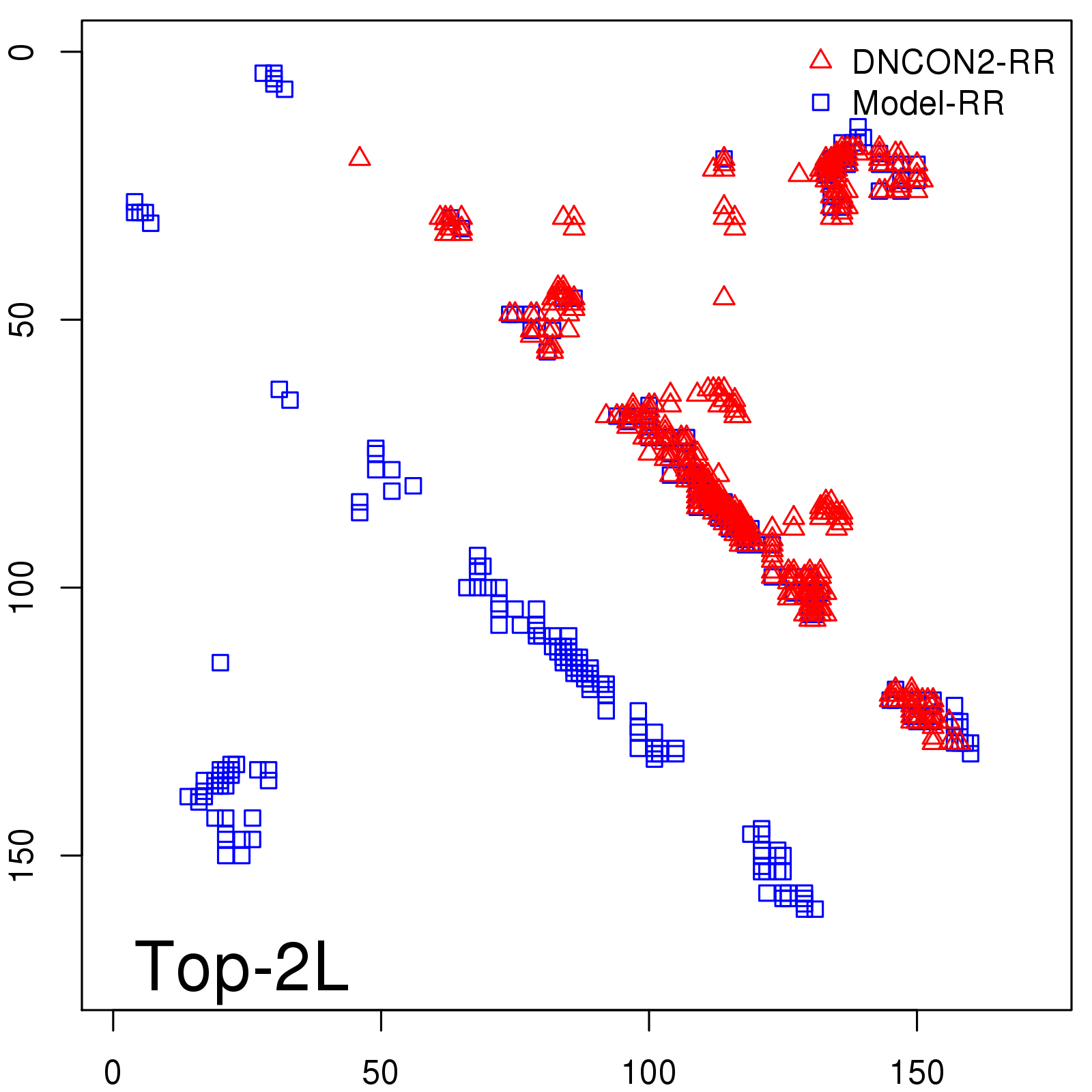
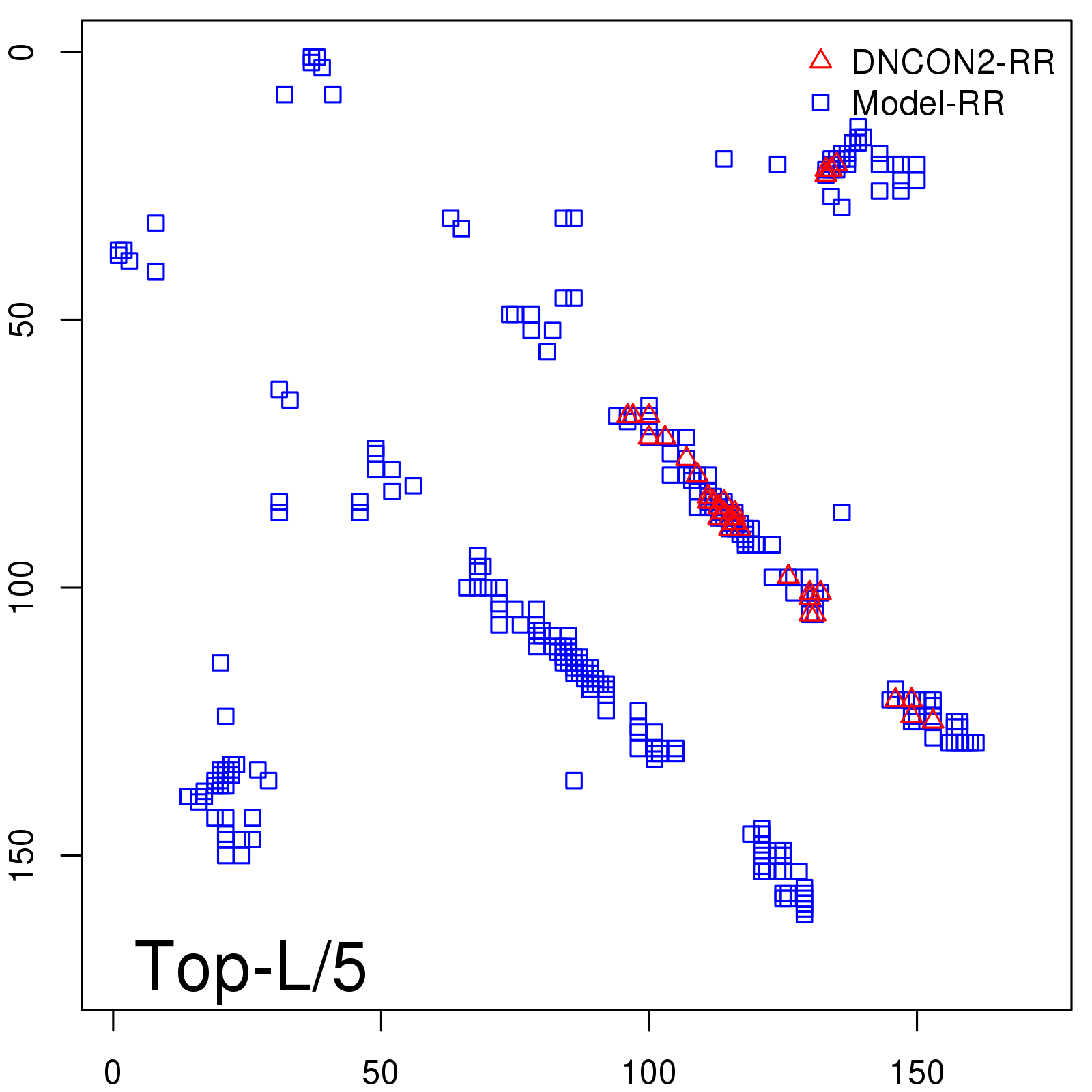
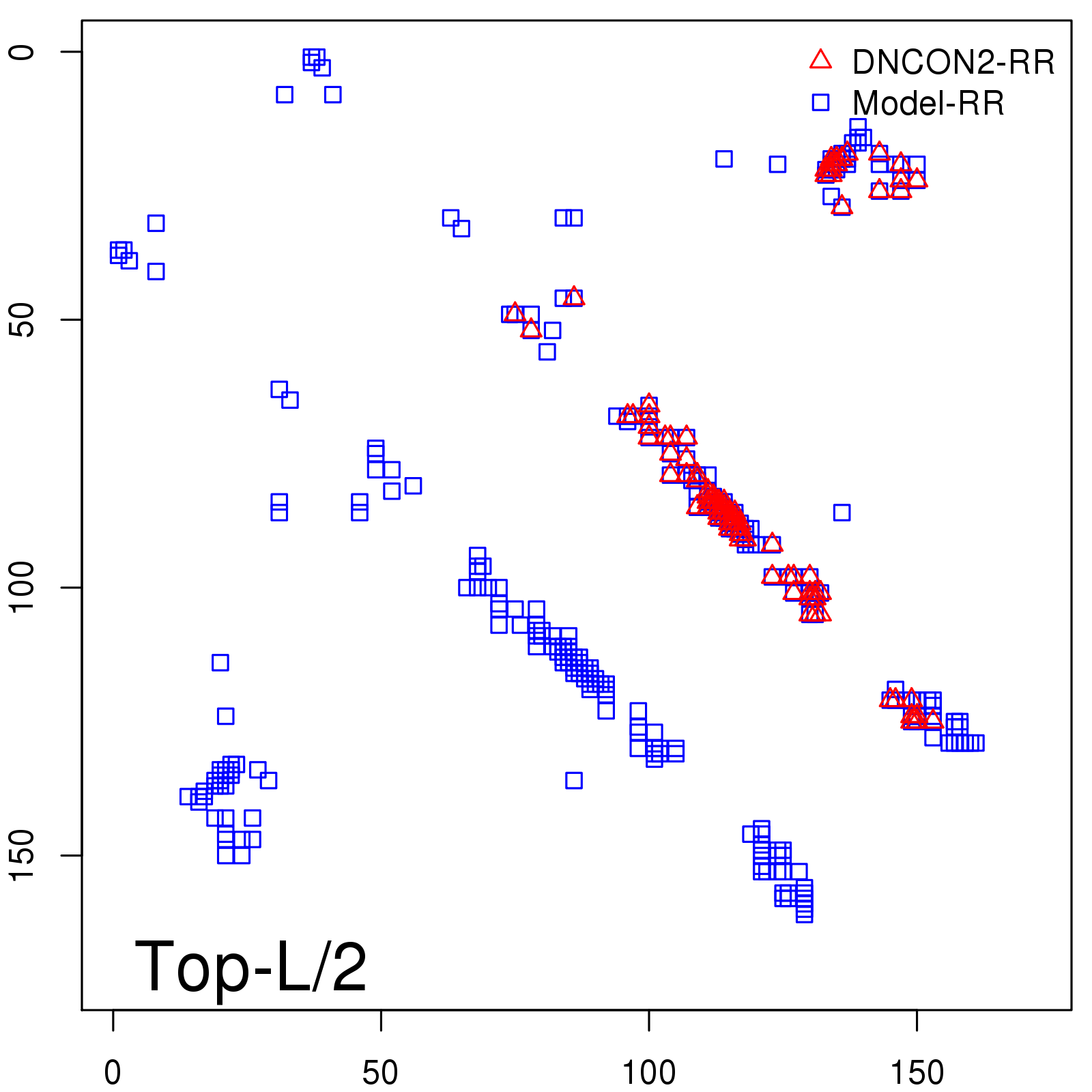
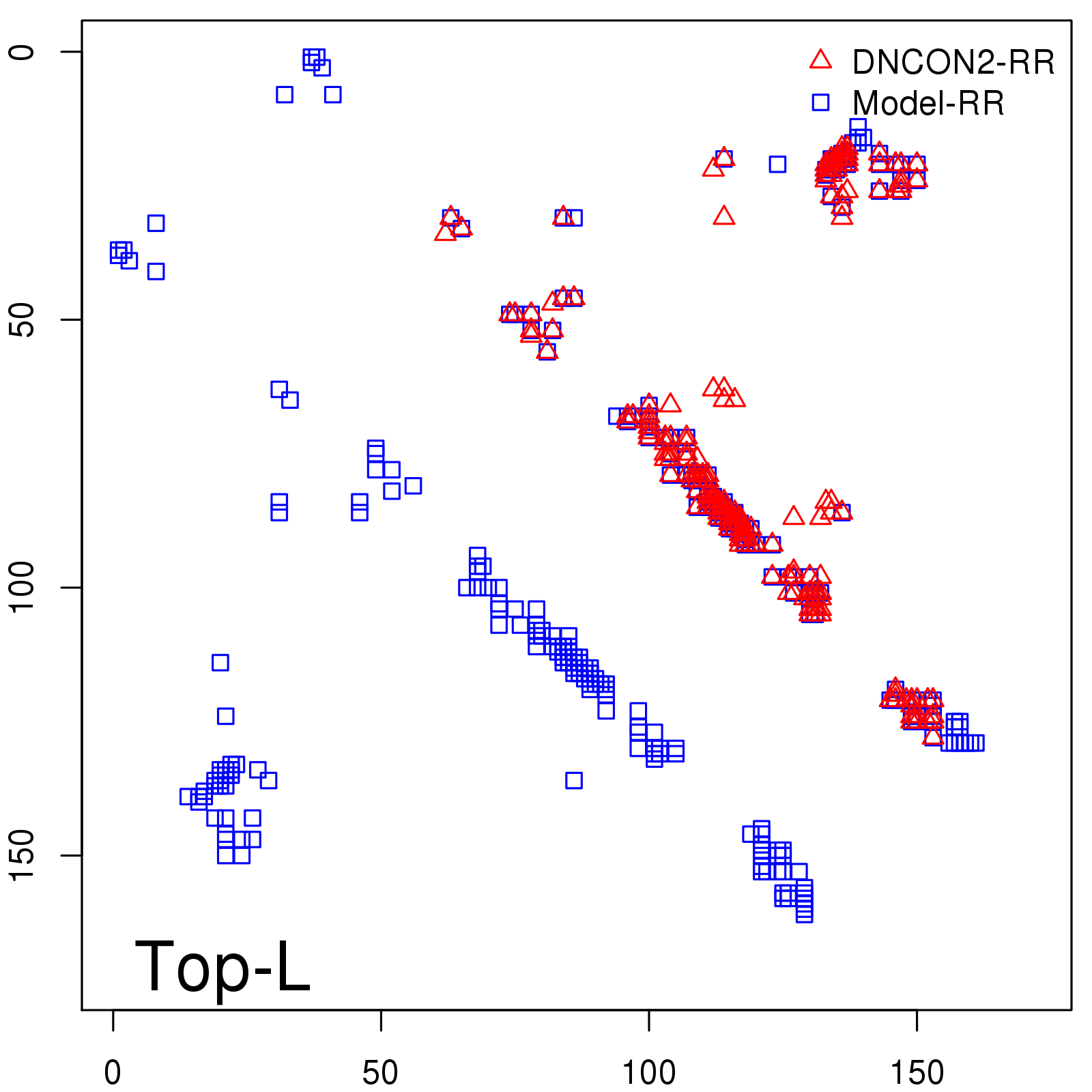
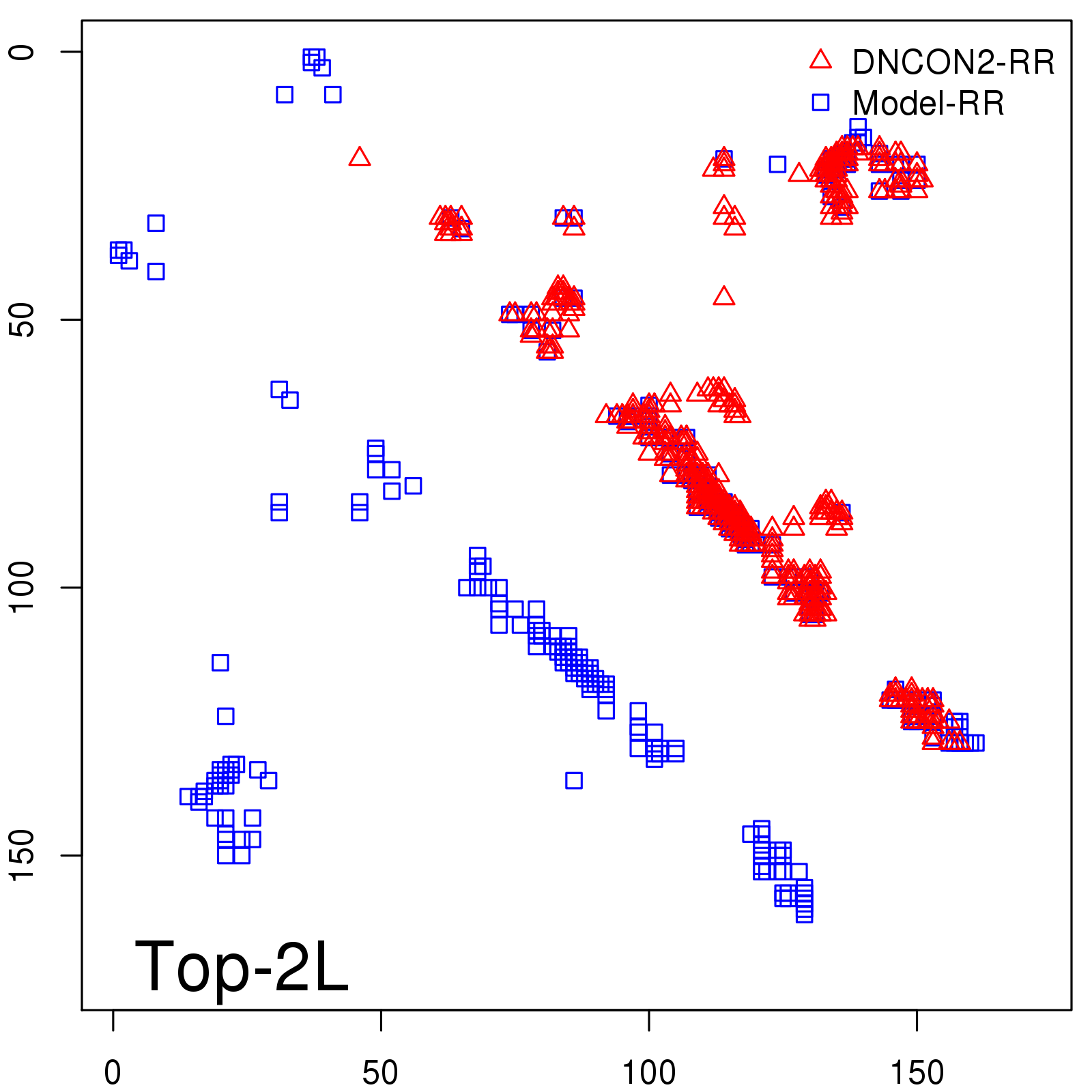
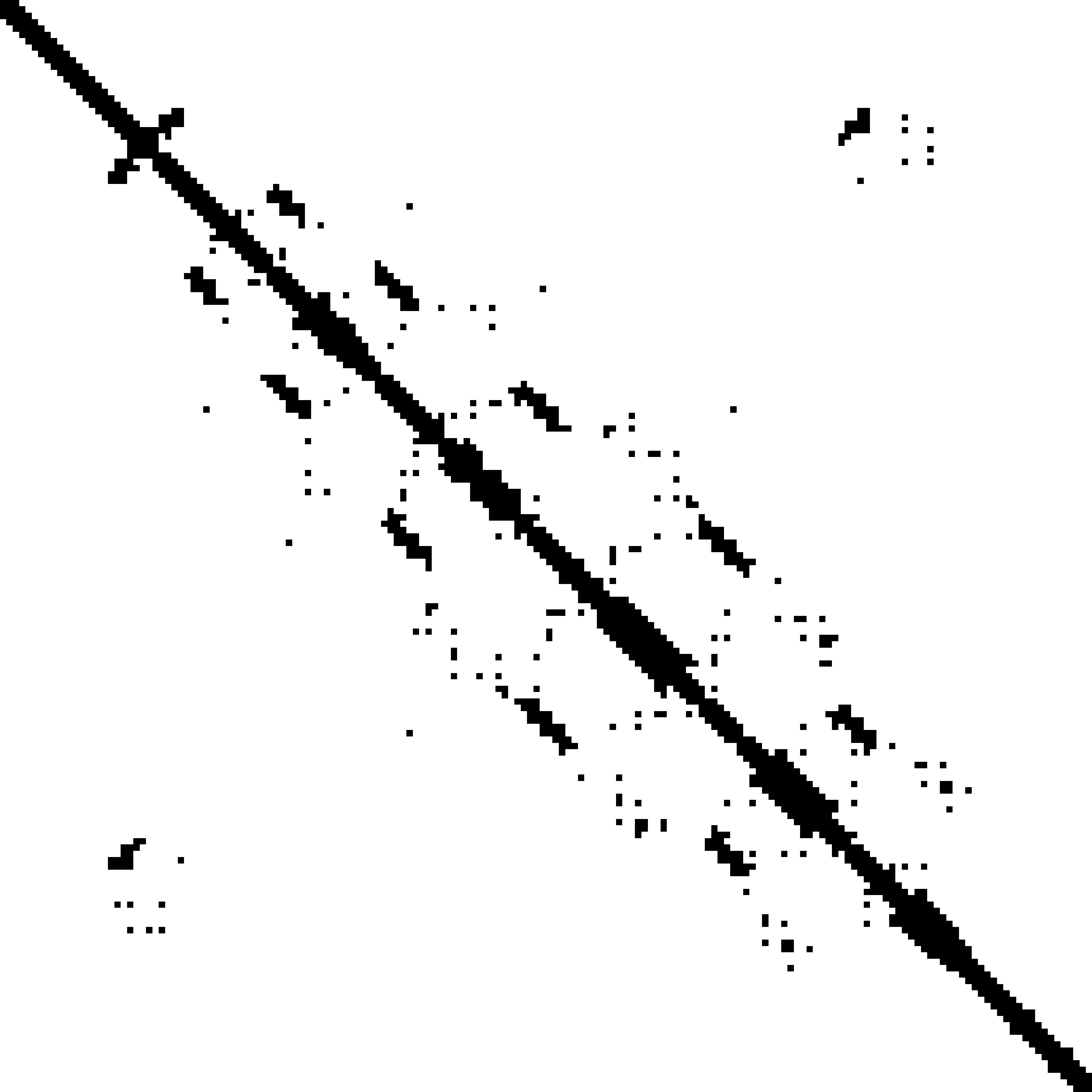
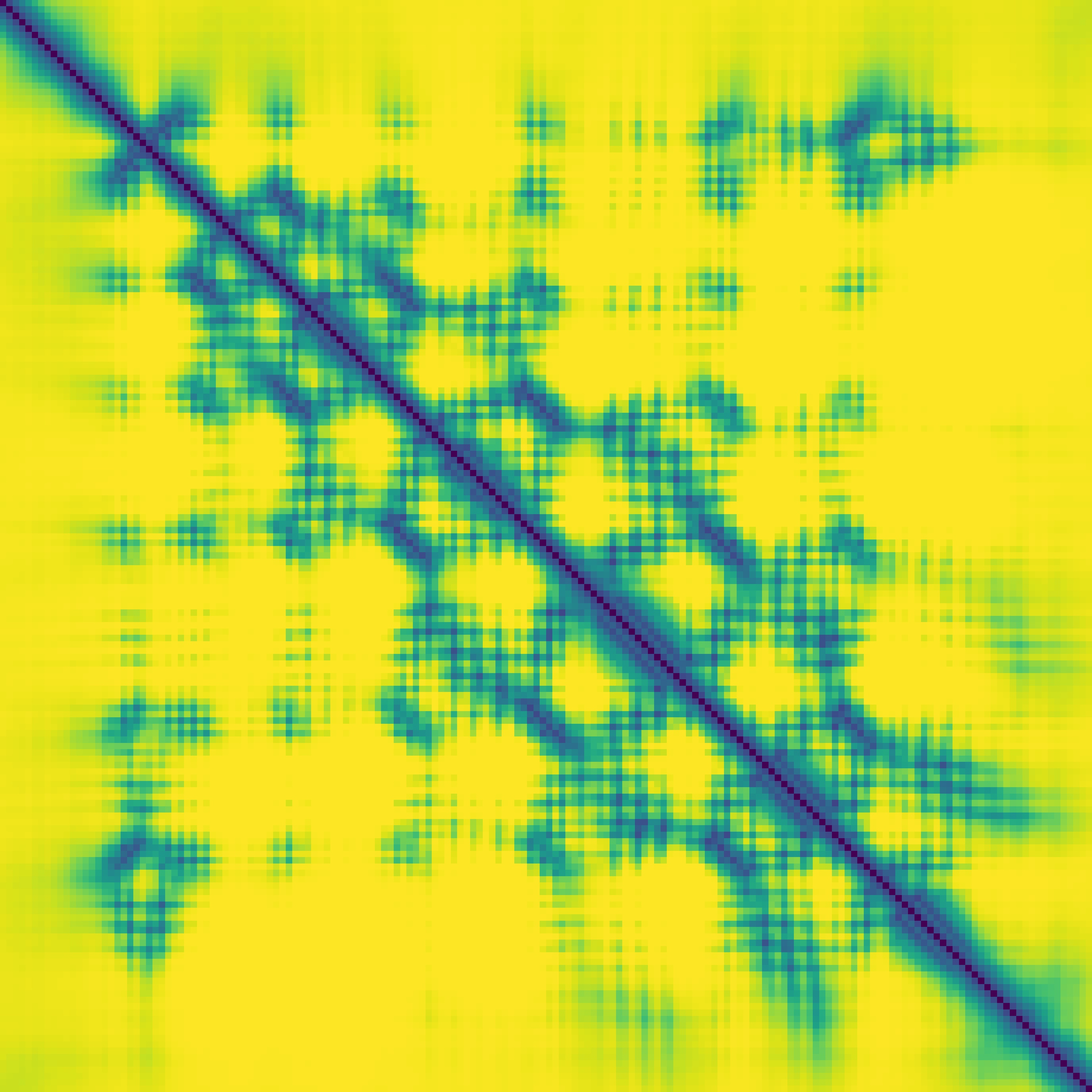
Predicted contact map and distance map
|
Predicted distance map 
|
|
Predicted contact map 
|
Probability to Precision
| Long-Range |
Average probability |
Predicted precision |
| TopL/5 |
0.92 |
91.08 |
| TopL/2 |
0.78 |
81.46 |
Note: The linear model (Average probability vs Predicted precision) was trained on CASP14 dataset
|
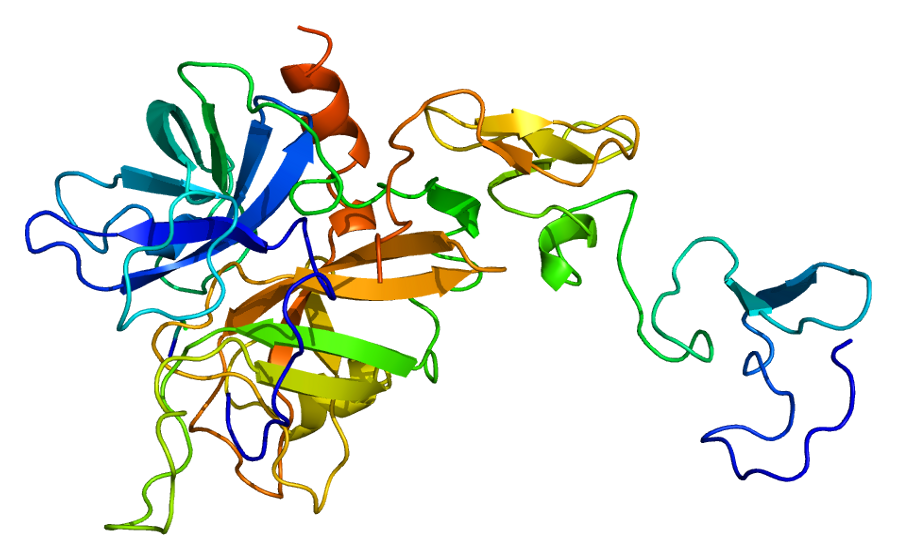
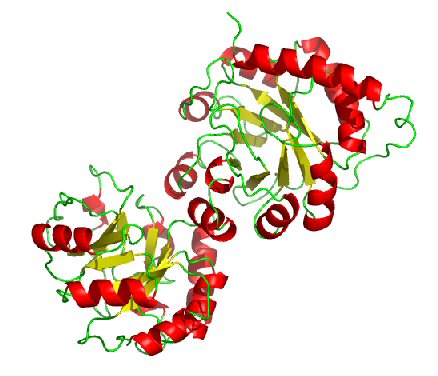
Predicted Top 1 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
100.00 |
| TopL/2 |
94.19 |
| TopL |
66.28 |
| Top2L |
34.59 |
| Alignment |
Number |
| N |
1258 |
| Neff |
390 |
|
Model comparsion
| Model |
TM-score |
| Model 2 |
0.9233 |
| Model 3 |
0.9457 |
| Model 4 |
0.9120 |
| Model 5 |
0.9308 |
| Average |
0.92795 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5zf4 |
0.70569 |
| 3zws |
0.70501 |
| 4jtu |
0.70473 |
| 5zf7 |
0.70465 |
| 6fmd |
0.70460 |
|
Note: The model is built by ab initio method, no significant templates are found!
|
Predicted Top 2 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
100.00 |
| TopL/2 |
95.35 |
| TopL |
65.70 |
| Top2L |
34.88 |
| Alignment |
Number |
| N |
1258 |
| Neff |
390 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.9233 |
| Model 3 |
0.9188 |
| Model 4 |
0.9030 |
| Model 5 |
0.9188 |
| Average |
0.91597 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 2prl |
0.72438 |
| 2fpt |
0.72419 |
| 1d3g |
0.72389 |
| 5hqe |
0.72326 |
| 5zf4 |
0.72322 |
|
Note: The model is built by ab initio method, no significant templates are found!
|
Predicted Top 3 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
100.00 |
| TopL/2 |
94.19 |
| TopL |
66.28 |
| Top2L |
36.05 |
| Alignment |
Number |
| N |
1258 |
| Neff |
390 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.9457 |
| Model 2 |
0.9188 |
| Model 4 |
0.9022 |
| Model 5 |
0.9470 |
| Average |
0.92843 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5k9d |
0.70470 |
| 2prl |
0.70426 |
| 5hin |
0.70407 |
| 5zf4 |
0.70386 |
| 3zws |
0.70384 |
|
Note: The model is built by ab initio method, no significant templates are found!
|
Predicted Top 4 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
100.00 |
| TopL/2 |
93.02 |
| TopL |
63.37 |
| Top2L |
33.14 |
| Alignment |
Number |
| N |
1258 |
| Neff |
390 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.9120 |
| Model 2 |
0.9030 |
| Model 3 |
0.9022 |
| Model 5 |
0.9191 |
| Average |
0.90908 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5k9d |
0.69992 |
| 5hqe |
0.69953 |
| 6j3c |
0.69931 |
| 5hin |
0.69919 |
| 5zf4 |
0.69897 |
|
Note: The model is built by ab initio method, no significant templates are found!
|
Predicted Top 5 Tertiary structure
|
|
|
| Long-Range |
Precision |
| TopL/5 |
100.00 |
| TopL/2 |
96.51 |
| TopL |
69.19 |
| Top2L |
36.34 |
| Alignment |
Number |
| N |
1258 |
| Neff |
390 |
|
Model comparsion
| Model |
TM-score |
| Model 1 |
0.9308 |
| Model 2 |
0.9188 |
| Model 3 |
0.9470 |
| Model 4 |
0.9191 |
| Average |
0.92893 |
Radius Gyration
|
Similar pdbs
| PDB |
TM-score |
| 5k9d |
0.70716 |
| 5zf4 |
0.70678 |
| 5hqe |
0.70675 |
| 5hin |
0.70659 |
| 6idj |
0.70633 |
|
Note: The model is built by ab initio method, no significant templates are found!
|
References
[1] Hou, J., Wu, T., Cao, R., & Cheng, J. (2019). Protein tertiary structure modeling driven by deep learning and contact distance prediction in CASP13. bioRxiv, 552422.
[2] Li, J., Deng, X., Eickholt, J., & Cheng, J. (2013). Designing and benchmarking the MULTICOM protein structure prediction system. BMC structural biology, 13(1), 2.
[3] Cheng, J., Li, J., Wang, Z., Eickholt, J., & Deng, X. (2012). The MULTICOM toolbox for protein structure prediction. BMC bioinformatics, 13(1), 65.
[4] Wang, Z., Eickholt, J., & Cheng, J. (2010). MULTICOM: a multi-level combination approach to protein structure prediction and its assessments in CASP8. Bioinformatics, 26(7), 882-888.
 )
) )
)

 )
) )
) )
) )
) )
)